Alberta Health Services genetic testing 2019 overview
Dr.M.Raszek
Alberta enters the genomic world
As we usher in this new era of DNA testing that is privately available to any consumer worldwide, perhaps we should first focus on the genetic testing that is publicly available in Alberta. While Merogenomics will always prefer and promote full genome sequencing for its completeness of DNA sequencing and the benefits that come with that, it is also important to be aware what is comparatively available to the public through their healthcare system.
The world of genetics and genomics is vast both in private access to medical genetics as well as public. But how vast is it here in Alberta? We catalogued all of the different tests available in the Alberta Health Services Genetics and Genomics program and were not at all disappointed with its size and complexity although some limitations can always be pointed out. One right off the top would be the obvious lack of uniformity in the information being listed with some procedures providing wonderful details, while some are missing even basic elements. We will touch upon that later, but that should be the first thing remedied. In the medical world where doctors are supposed to provide enough details to obtain accurate, informed consent from their patients then these are the very pertinent and important details that doctors need to be able to access easily.
Our other small complaint is that the information is not on an easy to navigate platform, but we do applaud what has been achieved thus far and hope that this is a great starting point to expand this information in the quickly changing medical world where more and more mainstream doctors are being expected to understand the potential and usage of DNA testing.
Overall, we counted 225 available tests. In our count, some procedures were consolidated into a single test if they were separated merely due to being performed on different tissue types. In theory, Alberta Health could provide a limitless number of tests, as doctors could possibly test for unique mutations of any kind if there is a documented history of it with strong evidence of pathogenicity associated with the mutation. However, the trick is to find that information in the first place. Usually, in the Alberta healthcare system, a referral would have to be made based on a clearly evident family history of a genetic condition, which could only then be attempted to be uncovered or else alternatively, the patient would have to rely on private clinical genetic testing to obtain the diagnostic evidence. That is the purpose of Merogenomics, to aid with access to clinical DNA testingthat otherwise might not be publicly available. This is the reason we chose to spend all this time in an analysis of the Alberta Health Services Genetics and Genomics tests, to provide an educated opinion on where it stands.
With the 225 tests that are listed, there are many ways of categorizing the tests. Some of these tests are for one specific condition; others are broader in nature and could be analyzed for a multitude of biological effects. In addition to the 225 tests, we also counted 6 tests that were specifically listed as being performed out of province so we did not count these, especially since details on these were sparse. In addition there were also 10 procedures that we categorized as supporting research tests – the aim of which is to collect additional information that might aid with utility or the interpretation of the results of some of the diagnostic genetic tests. An example of that would be a test to assign a family relationship between different samples, or a procedure to prepare the sample for subsequent testing (such as establishing a cell culture).
Genetic vs. metabolic tests
The first obvious division of the 225 tests: we established 164 to be of genetic nature (related to using DNA or RNA to obtain clinical information), and 60 were tests that assessed other biomarkers – most typically metabolites or other sample constituents such as proteins (which we collectively refer to as tests of metabolic nature). One test was pharmacokinetic in nature, or basically assessing what happens to a drug once it is in our blood system. In the past, metabolic testing was the easiest means of testing for heritable diseases but while these tests do not gauge the genetic makeup directly, all metabolic tests are still used for assessing heritable conditions, hence why they are listed by Alberta Health System under the Genetics and Genomics category. Since our focus is genetic testing, we will not focus on metabolic testing in much detail but rather we will dive into the specifics of DNA testing.
Another simple way to break down the 255 tests is to go by those that assess heritable conditions (conditions that can be passed on from one generation to another and we counted 189 tests), versus conditions that are of somatic origin (they arise due to DNA mutations after birth and therefore are not inherited – with 30 tests counted). Lastly, there is when a test meets both conditions (6 tests where the procedure could be used to study either a heritable condition or of a somatic nature).
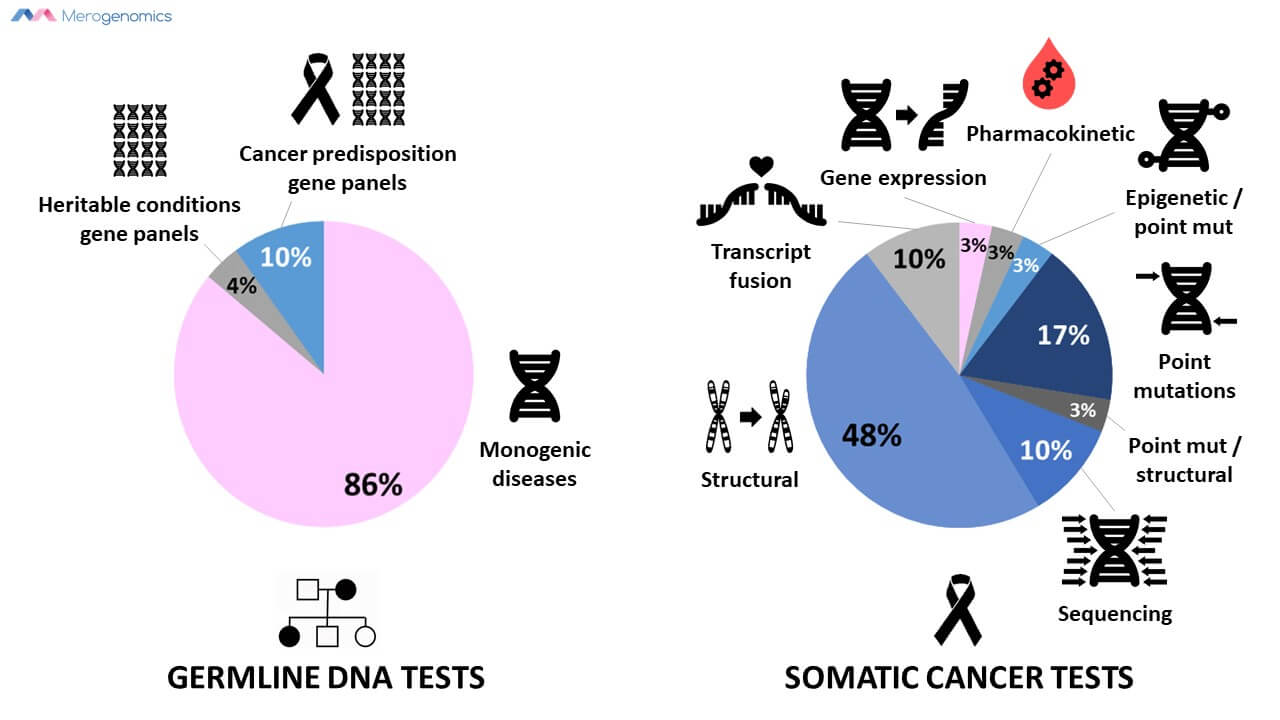
Of the somatic tests, all but one are dedicated to cancer analysis. Of the heritable conditions, 14 tests are dedicated to assessment of hereditary cancer predisposition, for a total of 49 tests that are dedicated to cancer genetic analysis. However, this might be an overestimate as some of the test descriptions do not clearly indicate if the procedure is used specifically for somatic or inherited mutation analyses. If the test information did not specify but could be used for both, it was placed in the “both” category.
Interestingly, during our cursory review of what type of conditions were investigated, we could ascertain that at least 17 of heritable conditions could also be of sporadic nature. This means that the condition might not actually have been inherited from afflicted parents. Rather, the DNA mutation that led to the disease spontaneously occurred in either the sperm or the egg that was used to produce the baby. In theory, the mutation could also occur after fertilization in the zygote. Such a resulting condition is heritable itself and thus could also then be passed onto the subsequent generation (although at times this is very unlikely due to the reduced reproductive success of afflicted individuals because of the high morbidity associated with the condition or an ensuing severe intellectual disability).
A good example would be Williams-Beuren syndrome which arises due to a whole chunk of a chromosome 7 going missing in one of the parents’ gametes with more than couple dozen genes gone along with it. When such a large segment of DNA is affected, it is referred to as a structural DNA mutation. As a consequence, multi-organ developmental issues arise, as well as cardiovascular problems, learning difficulties and distinctive physiological and behavioural features are produced. Such individuals can still pass on the condition to their children.
The genetic testing of heritable conditions can be further subcategorized into testing for mutations that affect the germline (alterations in the DNA of sperm or eggs) versus those affecting the mitochondrial DNA. Mitochondrial DNA is unique in that it is only inherited from the mother. Mitochondria are like tiny cells within cells and they are like miniature factories responsible for the production of ATP, dubbed “the energy molecule”. Only one Alberta Health Services genetic test is dedicated to the assessment of a mitochondrial conditions. The remaining 128 tests are all related to germline mutations.
Types of genetic information
Yet another way to categorize the tests is by the type of genetic information they analyze: epigenetics, gene expression, point mutations, DNA sequence data, structural information and finally RNA transcript fusion. Sequencing was further subcategorized as partial (where only a segment of the gene of interest is investigated for the DNA code information) or full sequencing (where all the exons of the gene are investigated). Only one test investigates introns (DNA that is present in between exons of the gene. It is the exons that are actually transcribed to produce the RNA, the introns can play important regulatory roles).
Most often multiple sources of genetic information are investigated in a single test, with 65 tests employing such an approach. In second place with 54 tests is the analysis of structural information, most typically either large deletions within the DNA or duplications of DNA segments. Some tests are specifically designed to look for more unusual complications such as inversions, where a segment of DNA is switched around. Third in line with 27 tests, are tests dedicated to assessing point mutations, or changes in a single DNA nucleotide position, also referred to as SNPs.
These are extremely targeted tests that are very limited in nature, meant to only diagnose that single change in the DNA position. If an equally damaging change in the DNA were to happen right beside it, such tests would miss this information. Such tests are cheap and easy to execute but they can miss a lot of potentially important information that could be used in helping diagnosing the patients. For these reasons, the broader sequencing of DNA is always better as it allows the process of discovery for new mutations. 14 tests are dedicated solely to sequencing the DNA, with only two of these being full sequencing. Three tests look at RNA transcript fusions in certain cancers, and one test assessed gene expression in cancer cell lines for prognostic purposes of cancer recurrence.
The good news is that of the mixed approach tests, 52 tests are dedicated at analyzing the complete gene sequence along with independent technology used to look for potential structural changes to the DNA (one of the tests also assessed the epigenetic information). This is the most comprehensive way to actually analyze patient genetic information for clinical purposes and we are glad to see that this approach is taking a foothold in our province. Unless you are doing full genome sequencing – which has certain advantages for being able to study structural mutations without the need of use of secondary technology – sequencing of select genes ideally should always be complemented by additional testing for structural changes in order to obtain the most complete genetic information related to a clinical condition possible. Some conditions are more prone to arise due to structural DNA alterations than to point mutations but every time one or the other approach is not utilized, there is a risk of missing important diagnostic information.
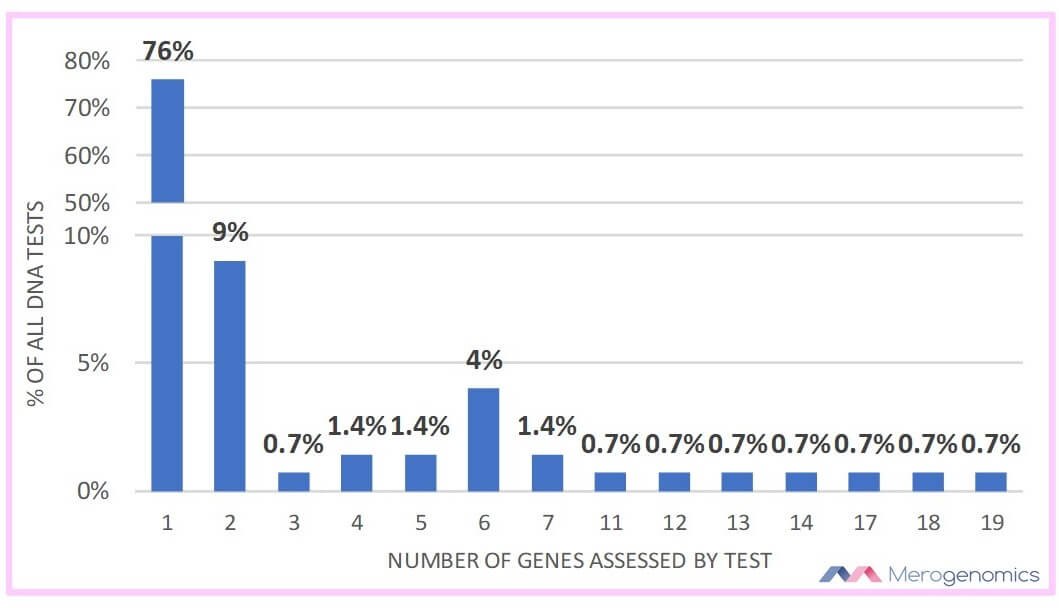
However, while that might sound all good, the reality is that the majority of these genetic tests that employ full sequencing and structural mutation analysis actually assess very limited information over all as the majority of these tests are dedicated to only a single gene evaluation. 138 of the tests list which genes are being studied and 76% of these are tests for just a single gene. While it can make sense when probing for a specific monogenic condition, the reality is that this is like trying to hit a bull’s-eye each time with only one throw. There are now nearly 6500 disease traits or symptoms that can be linked to molecular causes, which includes at least 4000 genes. With so many genetic conditions that have overlapping traits, the current clinical trend is steering towards much more comprehensive genetic analyses to increase the likelihood of diagnosis. One way is by pooling genes of interest into groups to assess the related condition which is referred to as a gene panel and this is rapidly becoming standard in medical genetics. If we count a panel as including minimum of three genes, then in total the Alberta Health Services system uses 20 tests with gene panels, with the largest panel including 19 genes.
Cancer predisposition testing
14 of these panels are for hereditary cancer predisposition which includes all the panels assessing seven or more genes. It is definitely exciting to see that Alberta Health Services is following suit in expanding the number of genes explored in cancer predisposition as more and more genes are being linked with this group of diseases. However, when it comes to actual cancer biopsy analysis for genetic information, the majority of the tests (62%) are dedicated to analyze broad structural genomic changes. Only 9 genes in cancer biopsies are analyzed for any specific sequence information. These include BRAF, EGFR, JAK2, KIT, KRAS, MLH1, MPL, NRAS and PDGFRA genes. Of these, only three are actually analyzed by sequencing, the remainder are assessed for specific point mutations only. This is a far cry from the type of analyses that can be performed, with full genome analysis of the tumor along with tumor transcriptome (sequencing of all of the RNA isolated from the tumor sample). Many medical centers around the world are already employing this approach, with a notable Canadian example of cancer medical genomics being found in Vancouver. Unfortunately, this procedure is very expensive, and therefore its public healthcare utility will be forthcoming very slowly.
Genetic testing technologies
How the above information is obtained depends on the type of technology used to collect the data and a very broad array of different methods are employed in the Alberta Health Services genetics and genomics tests. As the name of the program has been advanced to include genomics, so has the adoption of technology used in the program. In fact, next generation sequencing (NGS) is the most frequently employed tool in all of the genetic tests, whether alone or in combination with others. However, as previously alluded to, 60% of the time, this powerful technology is used to assess only single genes. One criticism that has to be made is that frequency with which the technology is used is not clearly specified, and thus our analysis is strictly based on the information that has been presented. About 25% of the listed tests do not clarify the methodology employed and we hope this will be remedied soon. For example, the third most commonly listed method is “Targeted Sequence Analysis” but this term could be describing a multiple of different potential approaches. Since we have no way of knowing, we placed it in its own category for now.
The second most frequently used technology is polymerase chain reaction (PCR), reflecting the targeted approach in genetic diagnosing currently used by the Alberta Health. Here too it is possible that the technology would be subcategorised into further specialized PCR methods that might not have been listed properly. The fourth most common technology is multiplex ligation-dependent probe amplification (MLPA), reflecting the fact of how frequently structural DNA information is assessed along with the DNA sequence. Even more encompassing for this purpose is the use of comparative genomic hybridization array (Array CGH), which ranks in at number seven. The use of this technology along with NGS is one of the primary gold standards now employed in medical genetics. Among all the genetic tests listed by the Alberta Health directories, these two technologies are used in total of 10 tests. This includes only 6 gene panels.
Biological samples accepted for genetic testing
The wide array of technologies used is also reflected by the wide array of samples accepted for testing purposes. Different samples are usually used for genetic testing versus metabolic testing with the exception of blood, which is accepted for use in certain tests among both categories. Within genetic testing, blood is the most preferred option. The majority of tests that utilize blood as a sample source also accept amniotic fluid, chorionic villi, extracted DNA or fetal tissues. In contrast, the saliva sample that is most frequently seen used in direct-to-consumer DNA testing is accepted in only one type of genetic test covered by public healthcare.
Alberta Health Services genetic testing locations
All tests are performed in either Edmonton or Calgary, with four locations participating in testing in each city. Overall, Calgary takes the crown for the highest number of tests on their menu with 153 tests and Edmonton performing 104 tests. Only 11 of these tests are actually delivered in both cities. The most tests by any facility is 90 tests by the Alberta Children's Hospital Molecular Diagnostics in Calgary, followed by 47 tests by the Alberta Children's Hospital Biochemical Genetics, also in Calgary. Edmonton comes in third place with 42 tests performed by the University of Alberta Hospital’s Molecular Diagnostics department.
Calgary also takes the crown by delivering 70% of the available gene panel tests. Edmonton on the other hand has the distinction of leading in terms of the most cancer tests with 64% of all cancer related tests delivered only in Edmonton with an additional 6% of cancer linked tests performed in both cities.
However, keep in mind that these rankings are just in terms of the number of different tests available (225), and not the actual volume of tests being performed.
Speed of Alberta Health genetic testing
In terms of turn around time (TAT), many tests provide a range of time for when the results could be available. Across all 225 tests, the average TAT is 47 days for the slower option and 41 days for the expedited results. Fastest test results can be available in a day but the slowest results for many of the DNA sequencing tests take up to 4 months. Average TAT for genetic tests is 57 days for regular delivery and 50 days for the expedited approach. Metabolic tests yield results much faster, on average being available in 19 days. The number of tests per facility with the TAT is listed in a table below.
| Location | Test count | TAT Avg | TAT single test | ||||||
| Slowest | Fastest | Slowest | Fastest | ||||||
| Alberta Children's Hospital Biochemical Genetics | 47 | 23 | 23 | 45 | 1 | ||||
| Alberta Children's Hospital Cytogenetics | 12 | 22 | 18 | 42 | 7 | ||||
| Alberta Children's Hospital Molecular Diagnostics | 82 | 72 | 62 | 112 | 42 | ||||
| Tom Baker Cancer Center Pharmacokinetics | 1 | 1 | 1 | 1 | 1 | ||||
| University of Alberta Hospital Molecular Diagnostics | 34 | 64 | 54 | 112 | 3 | ||||
| University of Alberta Hospital Molecular Pathology | 35 | 26 | 25 | 112 | 1 | ||||
| UAH or ACH Cytogenetics | 3 | 21 | 16 | 28 | 14 | ||||
| UAH or ACH Molecular Diagnostics | 8 | 86 | 86 | 112 | 14 | ||||
| University of Alberta Hospital Cytogenetics | 4 | 28 | 28 | 42 | 21 | ||||
| University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 20 | 8 | 8 | 14 | 4 | ||||
| Alberta Children's Hospital Cytogenetics all tests | 15 | 21 | 18 | 42 | 7 | ||||
| Alberta Children's Hospital Molecular Diagnostics all tests | 90 | 73 | 64 | 112 | 42 | ||||
| University of Alberta Hospital Cytogenetics all tests | 7 | 25 | 23 | 28 | 14 | ||||
| University of Alberta Hospital Molecular Diagnostics all tests | 42 | 69 | 60 | 112 | 3 | ||||
Alberta ethnicities’ genetic testing
Finally, the last assessment of publicly available genetic testing concerns which ethnicities are targeted by any of the tests specifically, and this outcome is definitely very Alberta specific. The one group of people targeted the most by genetic testing is the Hutterites, with 10 tests dedicated for that purpose. This is not surprising as Hutterites are an example of an isolated group that is more restricted in terms of genetic diversity and as a consequence any carrier mutations of pathogenic diseases are more likely to be present. This is to be expected in a group of people that is especially restricted in partner selection to a specific geographical area and comprised of a limited number of individuals. In such circumstances, genetic conditions will appear more frequently, and this is referred to as a founder effect, and the affected population can be termed the founder population. DNA mutations that otherwise would be so rare that they would almost never be witnessed in a wider population, can accumulate and become prominent. For these same reasons we also observe a greater prevalence of certain conditions among the Ashkenazi Jewish population, or the French Canadian population. The Hutterite population in prairie provinces has been extensively studied for its genetic conditions, and this fact has even led to the development of Hutterite-specific diagnostic DNA chip that can screen for 30 diseases.
While the prominence of Hutterite population representation was not a surprise, what was a surprise were lack of other ethnicities, more specifically, lack of French Canadian representation. Like the Hutterites, the French Canadians in Alberta would be expected to comprise of isolated groups that could also run the risk of allowing genetic conditions to persist, and this is one of the largest minorities in Alberta.
The take home message from our first analysis of the publicly available genetic testing in Alberta is that the program is certainly dynamic and clearly adopting new technologies as they become available and clinically verified, although much further progress can and should be expected in the years to come. Take for example, exome sequencing which assess all of the genes in the human genome, is not listed at all, not even as an option for extreme circumstances, although we do not doubt that it already must be happening to some degree in Alberta public healthcare. It is simply too powerful of a technology to save human lives to be ignored. Full genome sequencing on the other hand, probably not any time soon. Although we already alluded to use of this technology in cancer molecular profiling, another important diagnostic need for this technology is for people afflicted with conditions with no clear clinical diagnosis available, which is just as useful as exome sequencing, but even more comprehensive and accurate.
It will be exciting to watch this field evolve in our province.
The final component of this analysis is our attempt to provide access to all of the genetic testing information we were able to collect in a single table format. Our Albertan doctors will be encountering DNA testing more and more frequently in the future and genetic testing will not just belong to specialized medical departments anymore, but rather will be served and used by general doctors. Part of that vision requires continuous education on genetic testing by our medical establishment and developing a familiarity of use with these technologies in their every day practice. Currently there are many specialists that do not even think of DNA testing as a potential diagnostic tool and this will have to change in time as our grasp of genetics continues to grow, and we can see the benefits continue to be established, recognized, and utilized.
It is our goal to continue analyzing the exciting progress of genetic testing in Alberta in the years to come!
Please click on the letter to go to specific group of Alberta Health Services genetic tests:
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| 15q11-13 Duplication Syndrome | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics |
6 - 8 weeks | MLPA | Detect deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||||
| 1p19q Co-deletion by FISH for brain cancers | Investigation of oligodendrogliomas | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Tuesday | FISH | Slides are cut from paraffin block and are deparaffinized in xylene and dehydrated in ethanol. Specimen pretreatment is performed using SPoT-Light Tissue Pretreatment Kit (Life Technologies) and fluorescence in situ hybridization is performed using Vysis probes (Abbott Molecular Inc.) Vysis 1p36/1q25 and 19q13/19p13 FISH Probe Kit detects deletion of 1p36 and/or 19q13. In normal cells, there is an equal number of red and green signals (ratio of test probe to reference probe >0.8), but in abnormal cells the result is less red signals than green signals, indicating deletion of 1p or 19q (ratio of test probe to reference probe <=0.8). A ratio of test probe to reference probe <=0.8, and >15% of cells showing reference to target ratio of at least 2 is indicative of a deletion All appropriate negative and positive controls are used. An average of 100 interphase nuclei are examined for each probe set by two observers using an Olympus Provis Fluorescence microscope system on 1000X magnification |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin as well as six 4-micron baked (2 hours at 60°C) unstained charged slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
|||||||||||||
| 1q21.1 Deletion Syndrome | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Real-Time PCR | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| 22q11.2 Deletion Syndrome | DiGeorge/Velocardiofacial Syndrome (VCF) Shprintzen Syndrome Takao Syndrome CATCH22 |
University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Real-Time PCR | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| 3-Methylglutaconic & 3-Methylglutaric Acids | 3-Methylglutaconic and 3-methylglutaric acids can be modestly elevated in certain mitochondrial diseases, while secondary 3-methylglutaconic acidurias result in moderate to substantive elevations. Primary 3-methylglutaconic aciduria leads to gross elevations. These metabolites can also be elevated in plasma of patients with renal insufficiency, resulting in artefactual findings Primary 3MGC Disorders Detectable by elevated 3-methylglutaconic and 3-methylglutaric acids* -3-methylglutaconyl-CoA hydratase deficiency (3MGC type I; AUH) -3-hydroxy-3-methylglutaryl-CoA lyase deficiency Secondary 3MGC Disorders Detectable by elevated 3-methylglutaconic and 3-methylglutaric acids* -Barth syndrome (3MGC type II; TAZ) -Costeff Optic Atrophy syndrome (3MGC type III; OPA3) -MEGDEL syndrome (3MGC type IV; SERAC1) -Dilated Cardiomyopathy with Ataxia syndrome (3MGC type V; DNAJC19) -TMEM70 defect (TMEM70) -3-hydroxy-3-methylglutaryl-CoA lyase deficiency Other known causes of elevated 3-methylglutaconic and 3-methylglutaric acids* -CLPB (caseinolytic peptidase B) deficiency -Carbamoyl phosphate synthetase I deficiency -Some mitochondrial diseases -Some neuromuscular diseases -Other metabolic disorders (due to secondary mitochondrial dysfunction) |
3MGC-P, 3MGC-S, 3MGC-U | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Derivatization (i.e., silylation) using a gas chromatography-mass spectrometry (GC-MS) stable isotope dilution (SID) method with a deuterium-labeled internal standard | Urine 3-methylglutaconic and 3-methylglutaric acids can help to inform whether elevated plasma levels are due to a true 3-methylglutaconic aciduria or renal insufficiency, both of which can lead to a mild to moderate elevations of plasma 3-methylglutaconic and 3-methylglutaric acids. This test will be reflexively performed when elevated 3-methylglutaconic and/or 3-methylglutaric acids are found by Organic Acids, Semi-Quantitative (ACH) |
Urine | Collect a 5 to 10 mL (1.0 mL min) random urine sample in plain urine container and freeze | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary blood tube not refrigerated -Primary urine container not refrigerated or frozen -Aliquot tube received thawed -Tube type not written on aliquot tube label This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided. Urine 3-methylglutaconic and 3-methylglutaric acids should not be ordered in conjunction with urine organic acids, as these metabolites are also reported as part of that test, and this quantitative test will be reflexively performed on any urine sample with elevated levels detected, sample volume permitting |
|||||||||||||
| Blood | Collect 1 x 4.0 mL (0.75 mL plasma or serum min) sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST)" |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
||||||||||||||||||||
| 7-Dehydrocholesterol | Investigation of Smith-Lemli-Opitz syndrome, and the level of 7-dehydrocholesterol is reported along with the cholesterol level | 7DHC | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis Contact for priority turn-around |
Derivatization (i.e., silylation) using a gas chromatography-mass spectrometry (GC-MS) stable isotope dilution (SID) method with a deuterium-labeled internal standard | Elevated 7-dehydrocholesterol (7-DHC) is highly suggestive of a biochemical diagnosis of Smith-Lemli-Opitz (SLO). Mild elevation of 7-dehydrocholesterol can also be detected in patients with hypercholesterolemia and in patients on certain medications (e.g., haloperidol, aripiprazole, trazodone) | Blood | Collect 1 x 4.0 mL (0.25 mL plasma or serum min) sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary blood tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Acylcarnitine Profile | Investigation of mitochondrial fatty acid oxidation disorders and certain organic acidurias. This test may also be ordered to rule out the possibility of these disorders in a patient being assessed for the ketogenic diet Disorders Detectable by Acylcarnitine Analysis*: Fatty Acid Oxidation Disorders: -short-chain acyl-CoA dehydrogenase (SCAD) deficiency -medium-chain acyl-CoA dehydrogenase (MCAD) deficiency -medium/Short-chain 3-hydroxyacyl-CoA dehydrogenase (M/SCHAD) deficiency -medium-chain 3-ketoacyl-CoA thiolase (MCKAT) deficiency -dienoyl-CoA reductase deficiency -very long-chain acyl-CoA dehydrogenase (VLCAD) deficiency -long-chain 3-hydroxyacyl-CoA dehydrogenase (LCHAD) and trifunctional protein deficiencies -carnitine-acylcarnitine translocase (CACT) deficiency -carnitine palmitoyltransferase I (CPTI) and II (CPTII) deficiencies -electron transfer flavoprotein (ETF) and ETF-dehydrogenase deficiencies (a.k.a., multiple acyl-CoA dehydrogenase deficiency [MADD] or glutaric acidemia type II) Organic Acid Disorders: -glutaric acidurias -propionic Acidemia (PA) -methylmalonic Aciduria (MMA) -isovaleric Aciduria (IVA) -3-hydroxy-3-methylglutaryl-CoA lyase deficiency -3-methylglutaconyl-CoA hydratase deficiency -3-Methylcrotonyl-CoA carboxylase deficiency -isobutyryl-CoA dehydrogenase deficiency -2-methylbutyryl-CoA dehydrogenase deficiency -2-methyl-3-hydroxybutyryl-CoA dehydrogenase deficiency -beta-ketothiolase deficiency -malonic aciduria -ethylmalonic encephalopathy *Further confirmatory testing is required for some of these conditions, as nonspecific findings, isobaric species and overlapping patterns may prevent definitive diagnosis in specific cases This test may also be used to monitor known patients with certain mitochondrial fatty acid oxidation disorders or organic acidurias on diet-modified treatment" |
AC-P, AC-S | Alberta Children's Hospital Biochemical Genetics | 14 days Test performed on a weekly basis Contact for priority turn-around |
Derivatization (i.e., butylation) using electrospray ionization tandem mass spectrometry (ESI-MS/MS) with deuterium-labelled internal standards | Acylcarnitine analysis enables the diagnosis of specific disorders of fatty acid oxidation and certain organic acidurias, as relevant primary enzyme deficiencies result in the accumulation of specific acylcarnitines Free carnitine is not reported as part of this test; therefore Carnitine, Total and Free - Blood (ACH) should be ordered separately if there is suspicion of a systemic carnitine deficiency Urine organic acids will be reflexively performed on any existing sample if the results of the plasma acylcarnitine indicate the need for further investigation or metabolite discrimination. Metabolites reported: acetylcarnitine (C2), propionylcarnitine (C3), butyryl-/isobutyrylcarnitine (C4), tiglylcarnitine (C5:1), isovaleryl-/2-methylbutyryl-/pivalylcarnitine (C5), hexanoylcarnitine (C6), 3-hydroxyisovaleryl-/2-methyl-3-hydroxybutyrylcarnitine (C5-OH), octenoylcarnitine (C8:1), octanoylcarnitine (C8), malonylcarnitine (C3-DC), decadienoylcarnitine (C10:2), decenoylcarnitine (C10:1), decanoylcarnitine (C10), methylmalonyl-/succinylcarnitine (C4-DC), glutarylcarnitine (C5-DC), dodecenoylcarnitine (C12:1), dodecanoylcarnitine (C12), adipyl-/methylglutarylcarnitine (C6-DC), tetradecadienoylcarnitine (C14:2), tetradecenoylcarnitine (C14:1), myristoylcarnitine (C14), suberylcarnitine (C8-DC), 3-hydroxymyristoylcarnitine (C14-OH), palmitoleylcarnitine (C16:1), palmitoylcarnitine (C16), sebacylcarnitine (C10-DC), 3-hydroxypalmitoleylcarnitine (C16:1-OH), 3-hydroxypalmitoylcarnitine (C16), linoleylcarnitine (C18:2), oleylcarnitine (C18:1), stearoylcarnitine (C18), 3-hydroxylinoleylcarnitine (C18:2-OH), 3-hydroxyoleylcarnitine (C18:1-OH), 3-hydroxystearoylcarnitine (C18-OH), hexadecanedioylcarnitine (C16-DC) |
Blood | Collect 1 x 4.0 mL (0.25 mL plasma or serum min) sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary blood tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided |
||||||||||||
| Adenine Phosphoribosyl-transferase | consistent with adenine phosphoribosyltransferase deficiency | APRT | Alberta Children's Hospital Biochemical Genetics | N/A | Blood | Collect 1 x 4 mL whole blood in EDTA (lavender) tube | Must be a test specific dedicated tube (cannot share with any other test) Sample needs to arrive at the ACH Biochemical Genetics Lab within 24 hours of time of collection who will then forward it to an outside reference lab |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided This test should NOT be used as a primary screen for adenine phosphoribosyltransferase deficiency. Clinical findings of urolithiasis or crystalline nephropathy WITH evidence of elevated urine 2,8-dihydroxyadenine (2,8-DHA or DHA; using test Purine and Pyrimidine Profile - Urine) are required for testing to proceed |
||||||||||||||
| Adenosine Deaminase & Purine Nucleoside Phosphorylase | Investigation of erythrocyte adenosine deaminase (ADA) and purine nucleoside phosphorylase (PNP) deficiencies as the cause of immunodeficiency | ADA | Alberta Children's Hospital Biochemical Genetics | 21 days Test performed on a biweekly basis Contact for priority turn-around |
ADA and PNP enzyme activities are measured spectrophotometrically | Plasma ADA activity is measured in addition to erythrocyte activities to aid in the initial diagnosis of ADA, but is also used to titrate dosing in known ADA-deficient patients on PEG-ADA therapy The plasma ADA activity as part of this test should not be used for the purposes of diagnosing adenosine deaminase 2 deficiency (a.k.a., polyarteritis nodosa, Sneddon syndrome, cat eye syndrome chromosome region, candidate 1 deficiency) as it is has not be validated for that purpose |
Blood | Collect 2 x 4.0 mL (3 mL plasma or serum min, 1 x 4.0 mL for Calgary zone) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) | For non-Calgary collection sites: Tube #1 (whole blood sample) - Do NOT centrifuge. Do NOT aliquot. Do NOT freeze. Place whole blood sample on ice. Transport sample on ice (not frozen). (same for Calgary collection site) Tube #2 (plasma aliquot) - Centrifuge, transfer plasma to aliquot tube, write ""NaHep"" or ""LiHep"" on the aliquot tube label and freeze aliquot tube. Transport sample on dry ice to accompany Tube #1 |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Pleural or cardiac effusion, sputum or other specimen types for the express purpose of tuberculosis testing will NOT be accepted Samples will be rejected if: -Tube #1 (whole blood sample) not refrigerated -Tube #2 (plasma aliquot) received thawed -Tube type not written on Tube #2 (plasma aliquot) label -Sample type other than heparinized whole blood and plasma (e.g., pleural effusion) is submitted |
||||||||||||
| ALK by FISH | ALK | Reflex test when ALK by IHC results are equivocal | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Thursday | FISH | Slides are cut from the paraffin block and are deparaffinized in xylene and dehydrated in ethanol. Specimen pretreatment and fluorescence in situ hybridization (FISH) is performed using the Vysis Paraffin Pretreatment IV & Post-Hybridization Wash Buffer Kit (Abbott Molecular Inc.) and the FDA approved Vysis ALK Break Apart Probe Kit (Abbott Molecular Inc.) The LSI ALK Dual Color Break-Apart probe hybridizes to the band region 2p23. In normal cells, the two probes (green and orange) are fused or close together. In abnormal cells, with a break at 2p23, the two probes separate The negative cut off is established at < 15% of the cells showing an abnormal signal pattern according to the Vysis interpretation criteria. All appropriate negative and positive controls were used. An average of # interphase nuclei were examined independently by two observers using an Olympus Provis Fluorescence microscope system on 1000X magnification |
Paraffin-Embedded Tissue | One slide stained with Hematoxylin & Eosin as well as four 4-micron baked (2 hours at 60°C) unstained charged slides need to be submitted Aptex slides will also be accepted Please provide a copy of the corresponding pathology report |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
|||||||||||||
| Allopurinol Loading Test | Determination of ornithine transcarbamylase heterozygote status in a female patient suspected based on clinical findings, or partial ornithine transcarbamoylase deficient males who have had normal or borderline urinary orotic acid levels under normal conditions and/or normal molecular genetic testing | OR/OD | Alberta Children's Hospital Biochemical Genetics | 14 days Test performed on a weekly basis Contact for priority turn-around |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with deuterium-labelled internal standards | Metabolites reported: orotic acid, orotidine | Urine | Random urine collection container AND four 24 hour urine collection containers (no preservative) | PRE Allopurinol Load: Collect 5 - 10 mL random urine sample POST Allopurinol Load: Label the first 24-hour collection container with the start time and collect the total urine output for the next six hours into the container. Repeat three more times with each contained dedicated to 24-hour period Urine specimen MUST be kept FROZEN during the entire collection and transport process |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Specimens received thawed -Specimens not received in primary collection containers |
||||||||||||
| Angelman Syndrome | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | 15q11 methylation sensitive MLPA and UPD15 Microsatellites | 15q11 methylation sensitive MLPA: Detects copy number changes and determines methylation status. Does not distinguish between imprinting center pathogenic variants and UPD UPD15 microsatellites: Detects UPD15 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||||
| Alpha-Fucosidase | Investigation of Fucosidosis (a.k.a., alpha-fucosidase deficiency) - a lysosomal storage disorder | A-FUC-P | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Alpha-fucosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | Blood | Collect 1 x 4.0 mL (0.25 mL plasma min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) |
At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze It is recommended that the patient go to ACH OP Lab for specimen collection If multiple lysosomal enzymes are requested on plasma, a minimum of 0.25 mL plasma is required for each enzyme requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question, and/or by the results of an abnormal Oligosaccharides Screen, Urine (UAH) and evidence from additional send-out testing Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label -Serum is submitted |
|||||||||||||
| Alpha-Galactosidase | Investigation of Fabry disease in male patients based on clinical presentation | A-GAL-P | Alberta Children's Hospital Biochemical Genetics | 14 days Test performed on a biweekly basis Contact for priority turn-around |
Alpha-galactosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | The measurement of plasma alpha-galactosidase A activity is not a recommended diagnostic test for Fabry disease in females. In addition to a reported 30% of affected female patients showing normal enzyme activity, the Biochemical Genetics laboratory has identified several female patients with moderately decreased enzyme activity that were not subsequently found to have a detectable GLA mutation. Recommend referral to the ACH Inherited Metabolic Disorders Clinic for detailed investigation and/or application for out-of-province funding for GLA gene-based testing | Blood | Collect 1 x 4.0 mL (0.25 mL plasma min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) |
At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze It is recommended that the patient go to ACH OP Lab for specimen collection If multiple lysosomal enzymes are requested on plasma, a minimum of 0.25 mL plasma is required for each enzyme requested |
"Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label -Serum is submitted |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Alpha-Glucosidase, T-lymphocyte | Investigation of Pompe disease (a.k.a., alpha-glucosidase deficiency, acid maltase deficiency, glycogen storage disease type II, glycogenosis type II) - a lysosomal storage disorder | A-GLU-T | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Alpha-Glucosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | The establishment of an IL-2 stimulated T-lymphocyte culture and storage of cell pellets for the purpose of lysosomal enzyme testing Residual enzyme activity below 15% of the mean of the reference range cannot be reliably detected by this assay. Type and age of onset of symptoms as well as additional diagnostic testing may be needed to confirm severity of disease present |
Blood | Collect 1 x 4.0 mL (3 mL whole blood min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) |
Primary tube must remain at room temperature at all times All samples SHOULD arrive before 14:00h and have been collected NO LONGER than 48 hours prior to arrival It is recommended that patients go to ACH OP Lab for specimen collection The 3.0 mL of whole blood will be sufficient to test for multiple lysosomal enzymes in T-lymphocytes, if requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question Samples will be rejected if: -Primary tube not refrigerated -Any aliquot tube is received -Any sample other than whole blood is submitted |
||||||||||||
| Alpha-Mannosidase | Investigation of alpha-Mannosidosis (a.k.a., alpha-mannosidase deficiency) - a lysosomal storage disorder | A-MAN-P | "Alberta Children's Hospital Biochemical Genetics" | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Alpha-mannosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | Blood | Collect 1 x 4.0 mL (0.25 mL plasma min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) |
At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze It is recommended that the patient go to ACH OP Lab for specimen collection If multiple lysosomal enzymes are requested on plasma, a minimum of 0.25 mL plasma is required for each enzyme requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question, and/or by the results of an abnormal Oligosaccharides Screen, Urine (UAH: OLIGO) and evidence from additional send-out testing Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label -Serum is submitted" |
|||||||||||||
| Alpha-Thalassemia / Mental Retardation, X-linked | XNP | SUNQUEST: ATRX-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Sequence Analysis | Sequence analysis of exons 7,8 and 9 only | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Amino Acid Quantitation | Possible inborn errors of amino acid metabolism, especially nonketotic hyperglycinemia and serine biosynthesis defects CSF/plasma glycine ratio required for diagnosis of glycine encephalopathy (nonketotic hyperglycinemia (NKH)) |
SFAAQ | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 7 days Test performed Monday to Friday Contact for priority turn-around |
Cerebrospinal Fluid | Collect 1.0 mL (Minimum test volume 500 µL) Sterile plastic screw-cap vial for Cerebrospinal Fluid Freeze immediately |
This test should be ordered in conjunction with plasma amino acid quantitation. The specimens for both tests should be collected at the same time For accurate interpretation of the ratio, a plasma sample collected within 24 hrs of the CSF collection is required |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
||||||||||||||
| Investigation of disorders of amino acid metabolism with specific neurologic presentations (e.g., nonketotic hyperglycinemia, serine biosynthesis disorders, GABA transaminase deficiency) | AA-C | Alberta Children's Hospital Biochemical Genetics | 7 days Test performed on daily basis Contact for priority turn-around |
Amino acids are detected using an Amino Acid Analyzer with internal standard | Metabolites reported: phosphoserine, taurine, phosphoethanolamine, aspartic acid, hydroxyproline, threonine, serine, asparagine, glutamic acid, glutamine, sarcosine, alpha-aminoadipic acid, proline, glycine, alanine, citrulline, alpha-aminobutyric acid, valine, cystine, methionine, cystathionine, isoleucine, leucine, tyrosine, beta-alanine, phenylalanine, beta-aminoisobutyric acid, homocystine, gamma-aminobutyric acid, ethanolamine, hydroxylysine, ornithine, lysine, 1-methylhistidine, histidine, tryptophan, 3-methylhistidine, anserine, carnosine, arginine |
Cerebrospinal Fluid | Collect 0.25 mL Sterile plastic screw-cap vial for Cerebrospinal Fluid Preference is to use second CSF collection vial with no visible blood contamination Indicate specimen type on label Freeze |
This test should be ordered in conjunction with Amino Acids - Plasma (ACH) Both samples should be collected simultaneously |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided |
|||||||||||||
| Possible inborn errors of metabolism. May help in evaluation of endocrine disorders, liver diseases, muscle diseases, neoplastic diseases, neurological diseases, nutritional disturbances, renal failure, and burns | AAQ | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 7 days Test performed Monday to Friday Contact for priority turn-around |
Metabolites reported: taurine, aspartic acid, threonine, serine, asparagine, glutamic acid, glutamine, glycine, alanine, citrulline, alpha-amniobutyric acid, valine, cystine, methionine, isoleucine, leucine, tyrosine, phenylanine, tryptophan, ornithine, lysine, histidine, arginine |
Blood | Collect 1 x 4.5 mL (3 mL whole blood or 1 mL pediatric min) Lithium Heparin PST Gel (Light Green) Separate plasma. A minimum of 700uL of plasma must be submitted for testing Freeze plasma immediately |
Requires a minimum 4h fast;or just before the next feed for newborns. Physician test requests on non-fasted samples from known metabolic patients are also acceptable | Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Specimen Rejection Criteria: -Serum will not be accepted. -Grossly hemolyzed specimens -Plasma not received frozen |
||||||||||||||
| Investigation of disorders of amino acid metabolism This test may also be ordered to rule out the possibility of these disorders in a patient being assessed for the ketogenic diet This test may also be used to monitor known patients with certain amino acid disorders that are on diet-modified treatment Disorders Detectable by Amino Acid Analysis: -maple syrup urine disease (MSUD) -propionic Acidemia (PA) -isovaleric Acidemia (IVA) -methylmalonic Acidemia (MMA) -nonketotic Hyperglycinemia (NKH) -disorders of phenylalanine and tyrosine metabolism -hypermethioninemias -histidinemia -hypertryptophanemia -hyperlysinemias -aminoadipic aciduria -serine biosynthesis disorders -disorders of ornithine and proline metabolism -urea cycle disorders and inherited hyperammonemias- -mitochondrial energy metabolism disorders (i.e., primary lactic acidemias) -beta-alaninemia -cystathionase deficiency -glutamic acidemia -glutamine synthase deficiency |
AA-P | Alberta Children's Hospital Biochemical Genetics | 7 days Test performed on daily basis Contact for priority turn-around" |
Amino acids detected using Amino Acid Analyzer with internal standard | Metabolites reported: phosphoserine, taurine, phosphoethanolamine, aspartic acid, hydroxyproline, threonine, serine, asparagine, glutamic acid, glutamine, sarcosine, alpha-aminoadipic acid, proline, glycine, alanine, citrulline, alpha-aminobutyric acid, valine, cystine, methionine, cystathionine, isoleucine, leucine, tyrosine, beta-alanine, phenylalanine, beta-aminoisobutyric acid, homocystine, gamma-aminobutyric acid, ethanolamine, hydroxylysine, ornithine, lysine, 1-methylhistidine, histidine, tryptophan, 3-methylhistidine, anserine, carnosine, arginine This test is not a reliable method for detecting and quantitating homocystine, as levels can be altered depending on the length of time between sample collection and analysis, as well as the presence of an interfering peak that falsely suggests elevated homocystine. Recommend ordering Homocysteine, Total - Blood (ACH) or Homocysteine, Total - Urine (ACH) for evaluation of homocytine levels Similarly, various medications can lead to ninhydrin-positive peaks that interfere with the identification and quantitation of certain amino acids. It is recommend that all current medications being take at the time of specimen collection be provided in the reason for referral |
Blood | Collect 1 x 4.0 mL (0.25 mL min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) /td> |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) Physician test requests from known metabolic patients (non-fasting) are also acceptable At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label -Serum is submitted |
|||||||||||||
| Evaluating disorders of amino acid transport and a few select disorders of amino acid metabolism Amino Acids Screen, urine | UAAQ | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetic | 7 days Test performed Monday to Friday Contact for priority turn-around |
Urine | Collect 5 to 10 mL (1.0 mL min) random urine specimen in plain urine container Freeze immediately |
Test requires creatinine result for interpretation Urine amino acids screen is done initially and quantitation ordered if indicated |
Request using Chemistry Specialty Requisition Plasma amino acids is the recommended test for primary investigation. Include family history, clinical condition, diet and drug therapy information on the requistition |
|||||||||||||||
| Investigation of disorders of amino acid transport (e.g., cystinuria, Hartnup disorder) and selected disorders of amino acid metabolism that require confirmation and/or additional metabolite identification/discrimination Primary Disorders Detectable by Urine Amino Acid Analysis*: -cystinuria -Hartnup disorder -beta-alaninemia -disorders of pyrimidine metabolism -alpha-aminoadipic aciduria -dibasic aminoaciduria -HHH syndrome -lysinuric protein intolerance -argininosuccinic aciduria (argininosuccinate lyase deficiency) -dicarboxylic aminoaciduria -carnosinemia -saccharopinuria -cystathionase deficiency -hyperlysinemia -hyperornithinemia -hyperprolinemia -hydroxyprolinuria -hypophosphatasia -familial renal iminoglycinuria -tryptophanuria -Secondary Disorders Detectable by Urine Amino Acid Analysis*: -Fanconi syndrome -Vitamin D-dependent rickets *Further confirmatory testing is required for many of these conditions, as nonspecific findings and overlapping patterns may prevent definitive diagnosis in specific cases |
AA-U | Alberta Children's Hospital Biochemical Genetics | 14 days Test performed on daily basis Contact for priority turn-around |
Amino acids are detected using an Amino Acid Analyzer with internal standard | This test should not be used as a frontline test for disorders of amino acid metabolism; recommend Amino Acids - Plasma (ACH) as a frontline test Metabolites reported: phosphoserine, taurine, phosphoethanolamine, aspartic acid, hydroxyproline, threonine, serine, asparagine, glutamic acid, glutamine, sarcosine, alpha-aminoadipic acid, proline, glycine, alanine, citrulline, alpha-aminobutyric acid, valine, cystine, methionine, cystathionine, isoleucine, leucine, tyrosine, beta-alanine, phenylalanine, beta-aminoisobutyric acid, homocystine, gamma-aminobutyric acid, ethanolamine, hydroxylysine, ornithine, lysine, 1-methylhistidine, histidine, tryptophan, 3-methylhistidine, anserine, carnosine, arginine This test is not a reliable method for detecting and quantitating homocystine, as levels can be altered depending on the length of time between sample collection and analysis, as well as the presence of an interfering peak that falsely suggests elevated homocystine. Recommend ordering Homocysteine, Total - Blood (ACH) or Homocysteine, Total - Urine (ACH) for evaluation of homocytine levels. Similarly, various medications can lead to ninhydrin-positive peaks that interfere with the identification and quantitation of certain amino acids. It is recommend that all current medications being take at the time of specimen collection be provided in the reason for referral |
Urine | Collect 5 to 10 mL (1.0 mL min) random urine specimen in plain urine container If sample is transferred to a secondary aliquot tube or container, write ""random urine"" on label Freeze immediately Alternate: Min 10 mL 24 hour urine collection container (NO ADDITIVE) Instruct patient to ensure to place 24 hour urine collection container into freezer immediately after each void throughout the 24 hour collection period |
24 hour Urine Collection: 1. Thaw 24hr urine specimen 2. Measure total volume 3. Enter TVOL on requisiton 4. Send 10 mL aliquot of urine; write ""24hr urine, Tvol #L"" on the label 5. Freeze aliquot This test has limited clinical utility for investigating disorders of amino acid metabolism, and therefore Amino Acids - Plasma (ACH) is recommended |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Aliquot received thawed -Sample type not written on aliquot tube label |
|||||||||||||
| Aminoglycoside-Induced Ototoxicity | MTRNR1 | SUNQUEST: MTRNR1-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Sequence Analysis | Detects mutation m.1555A>G | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Amyotrophic Lateral Sclerosis | SOD1, C9ORF72 | Lou Gehrigs Disease SOD1 |
SUNQUEST: ALS-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | PCR and sequence analysis | Sequence Analysis for SOD1: Full gene sequencing PCR for C9ORF72: Two PCR reactions to detect repeats in the 4 - 60 range |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: -Dx - Neurology -Presymptomatic - Genetics |
||||||||||||
| Arrhythmogenic Right Ventricular Dysplasia / Cardiomyopathy | TMEM43 | ARVC ARVD TMEM43 |
SUNQUEST: ARVC5-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Detects the c.1073C>T (NFLD mutation) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics and Cardiology |
||||||||||||
| Arylsulfatase A, T-lymphocyte | Investigation of Metachromatic Leukodystrophy (a.k.a., arylsulfatase A deficiency) - a lysosomal storage disorder Can also be used for the further investigation of multiply sulfatase deficiency based on initial laboratory investigations showing abnormal urine glycosaminoglycan (i.e., dermatan and heparan sulfate elevations) and reduced iduronate 2-sulfatase activity |
ASA | Alberta Children's Hospital Molecular Diagnostics | 45 days Test is performed on a monthly basis Contact for priority turn-around |
Arylsulfatase A activity is measured spectrophotometrically using a p-nitrocatechol sulphate-linked substrate | The establishment of an IL-2 stimulated T-lymphocyte culture and storage of cell pellets for the purposes of Lysosomal Enzyme Testing | Blood | Collect 1 x 4.0 mL (3 mL whole blood min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) Primary tube must remain at room temperature at all times |
Collected NO LONGER than 48 hours prior to arrival It is recommended that patients go to ACH OP Lab for specimen collection The 3.0 mL of whole blood will be sufficient to test for multiple lysosomal enzymes in T-lymphocytes, if requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question Samples will be rejected if: -Primary tube not kept at room temperature -Any aliquot tube is received. -Any sample other than whole blood is submitted |
||||||||||||
| Ataxia Telangiectasia | ATM | SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 16 weeks | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on an NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. aCGH: Deletion/duplication testing is performed at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics and Neurology |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| BAP1 Tumor Predisposition Syndrome | BAP1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
||||||||||||||
| Bardet-Biedl Syndrome | BBS1, BBS2, ARL6 (BBS3), BBS5, MKKS (BBS6), BBS10 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Targeted testing is available for 10 common pathogenic variants: BBS1, c.1169T>G (NM_024649.4) Newfoundland population: -BBS2, c.72C>G (NM_031885.2) -ARL6 (BBS3), c.505G>C (NM_001323513.1) -BBS5, c.522+3A>G (NM_152384.2) -MKKS (BBS6), c.281delT, c.429_434del6insTT and c.830T>C (NM_018848) -BBS10, c.271dup and c.952_957del (NM_024685) Hutterite population: -BBS2, c.472-2A>G (NM_031885) |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Beckwith-Wiedemann Syndrome | CDKN1C, locus 11p15 | BWS | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | 11p15 methylation sensitive MLPA, CDKN1C sanger sequencing | 11p15 methylation sensitive MLPA: Designed to detect abnormal methylation of the two imprinted domains located within the chromosome 11p15 BWS/RSS gene cluster as well as deletions and duplications within the 11p15 region. This assay will detect UPD11 CDKN1C sanger sequencing: Analysis of all coding regions of CDKN1C |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Beta-Galactosidase, T-lymphocyte | Investigation of beta-galactosidase deficiency leading to either Morquio B Syndrome (a.k.a., mucopolysaccharidosis type VI) or GM1 Gangliosidosis (a.k.a., Landing disease) - lysosomal storage disorders | B-GAL | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis Contact for priority turn-around |
Beta-Galactosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | This procedure involves the establishment of an IL-2 stimulated T-lymphocyte culture and storage of cell pellets for the purposes of lysosomal enzyme testing | Blood | Collect 1 x 4.0 mL (3 mL whole blood min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) Primary tube must remain at room temperature at all times |
Collected NO LONGER than 48 hours prior to arrival It is recommended that patients go to ACH OP Lab for specimen collection The 3.0 mL of whole blood will be sufficient to test for multiple lysosomal enzymes in T-lymphocytes, if requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not kept at room temperature -Any aliquot tube is received. -Any sample other than whole blood is submitted This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question |
||||||||||||
| Beta-Glucosidase, T-lymphocyte | Investigation of Gaucher Disease (a.k.a., acid beta-glucosidase deficiency, glucocerebrosidase deficiency) - a lysosomal storage disorder | B-GLU | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis Contact for priority turn-around |
beta-Glucosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | This procedure involves the establishment of an IL-2 stimulated T-lymphocyte culture and storage of cell pellets for the purposes of lysosomal enzyme testing | Blood | Collect 1 x 4.0 mL (3 mL whole blood min) sodium heparin tube (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) Primary tube must remain at room temperature at all times |
Collected NO LONGER than 48 hours prior to arrival It is recommended that patients go to ACH OP Lab for specimen collection The 3.0 mL of whole blood will be sufficient to test for multiple lysosomal enzymes in T-lymphocytes, if requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not kept at room temperature -Any aliquot tube is received. -Any sample other than whole blood is submitted This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question |
||||||||||||
| Beta-Mannosidase | Investigation of beta-Mannosidosis - a lysosomal storage disorder | B-MAN-P | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Beta-mannosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | Blood | "Collect 1 x 4.0 mL (0.25 mL plasma min) sodium heparin tube (dark green) Alternate: Lithium heparin (dark green - NOT PST) |
It is recommended that patients go to ACH OP Lab for specimen collection If multiple lysosomal enzymes are requested on plasma, a minimum of 0.25 mL plasma is required for each enzyme requested At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label -Serum is submitted This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question and/or by the results of an abnormal Oligosaccharides Screen, Urine (UAH) and evidence from additional send-out testing |
|||||||||||||
| Biotinidase | Diagnosis of biotinidase deficiency based on clinical presentation or as a confirmatory test for screen positive newborns | BIOT-S, BIOT-P | Alberta Children's Hospital Biochemical Genetics | 21 days Test is performed on a biweekly basis Contact for priority turn-around |
Biotinidase activity is measured based on the amount of lysine liberated from the biocytin substrate using an amino acid analyzer | Partial deficiency or carriers may have activity at the low end of the reference interval Prematurity can lead to decreased enzymatic activity within the first days to weeks of life, and chronic liver dysfunction can also lead to decreased activity |
Blood | Collect 1 x 4.0 mL (0.5 mL serum or plasma min) Non-additive (red top - NOT SST) Alternate: -Sodium heparin (dark green - NOT PST) -Lithium heparin (dark green - NOT PST) |
At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided |
||||||||||||
| Biotinidase Quantitation, plasma | Diagnosis of biotinidase deficiency | BTDQ | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed Monday to Friday |
Blood | Collect 1 x 2 mL (200 uL plasma or serum min) Lithium Heparin PST Gel (LIGHT GREEN) or SST Gel No Additive (GOLD) Spin, separate and freeze plasma (or serum) within one hour of collection |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
|||||||||||||||
| Birt-Hogg-Dube / Primary Spontaneous Pneumothorax | FLCN | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Oncology and Respirology. Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||||
| Bowen-Conradi Syndrome | EMG1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.257A>G (NM_006331) in the Hutterite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics and Neonatology |
||||||||||||||
| BRAF Mutation Analysis | BRAF | Investigation of colon (therapy and HNPCC) cancers and melanoma | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Monday and Wednesday (excluding statutory holidays) |
PCR followed by capillary electrophoresis | For Colorectal Cancer: DNA extraction from FFPE samples is performed using the QIAGEN QIAsymphony DSP DNA mini kit. Analysis for BRAF, KRAS, NRAS mutations is performed after PCR amplification using a single base extension technique (SNaPshot multiplex kit by Life Technologies) and capillary electrophoresis on the 3130 Genetic Analyzer. This multiplexed assay is capable of detecting mutations at BRAF codon 600 and KRAS/NRAS codons 12, 13, 61, 117, and 146 The estimated sensitivity of this assay is 10% For Melanoma: DNA extraction from FFPE samples is performed using the QIAGEN QIAsymphony DSP DNA mini kit. Analysis for BRAF mutations is performed after PCR amplification using a single base extension technique (SNaPshot multiplex kit by Life Technologies) and capillary electrophoresis on the 3130 Genetic Analyzer. This multiplexed assay is capable of detecting mutations at BRAF codon 600 The estimated sensitivity of this assay is 5% |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred. Alternatively, one slide stained with Hematoxylin & Eosin as well as five 10-micron non-baked unstained slides can be submitted Please provide a copy of the corresponding pathology report |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
|||||||||||||
| Breast Ovarian Cancer Panel, Core | BRCA1, BRCA2, CDH1, PTEN, STK11, TP53 | HBOC Cancer and Endocrine Next Generation Sequencing |
SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE CERNER: MDLBL-;APPEND TEST NAME AND SPECIMEN TYPE |
University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | For UAH, maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Gynecological Oncology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Breast Ovarian Cancer Panel, Extended | ATM, BRCA1, BRCA2, BRIP1, CDH1, CHEK2, NBN, PALB2, PTEN, RAD51C, RAD51D, STK11, TP53 | |||||||||||||||||||||
| Busulfan | Patients undergoing stem cell transplantation and on Busulfan therapy protocol | CERNER: L-BUSULFEOI, L-BUSULF1HR, L-BUSULF3HR, L-BUSULF5HR, L-BUSULF7HR | Tom Baker Cancer Center Pharmacokinetics | Same day Prior arrangement is required before ordering |
HPLC DAD | Blood | Collect 1 mL plasma Sodium heparin (dark green - NOT PST) for each timed collection post infusion : 5 minutes, 1 hour, 3 hours, 5 hours, 7 hours | Immediately after each specimen collection, centrifuge blood specimen at 10oC for 10 minutes Transfer plasma into a plastic aliquot tube Call Pharmacokinetics Laboratory 403-521-3442 in advance of collection Write Na Hep plasma on the aliquot tube label Freeze aliquot at -20oC |
Direct order through SCM Order Restrictions: Can be ordered only by Bone Marrow Transplant Unit Physician |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| CADASIL | NOTCH3 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Sanger sequencing of exons 2-6, 11 and 18 of the NOTCH3 gene (NM_000435) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||||
| Calreticulin | CalR | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up Monday, bi-weekly |
PCR followed by capillary electrophoresis | After DNA extraction, PCR amplification for CalR gene is performed in a single tube using fluorescently labeled primers. Amplification products are subjected to fragment analysis by capillary electrophoresis using ABI 3130 genetic analyzer. Results were analyzed by Genemapper v3.2 software. The analytical sensitivity of this assay is 5% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
|||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| Carney Complex | PRKAR1A | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
||||||||||||||
| Carnitine - Total and Free | Investigation of a systemic carnitine deficiency due to an underlying primary (i.e., carnitine uptake defect due to OCTN2 deficiency) or secondary (e.g., MCAD deficiency, organic acidurias) cause This test can also be used to evaluate the potential of a secondary carnitine deficiency due to nutritional inadequacy (e.g., vegan diet, total parenteral nutrition), increased metabolic demand (e.g., ketogenic diet), reduced intestinal absorption (e.g., short bowel syndrome) or increased loss (e.g., Fanconi syndrome, hemodialysis) This test can also be used to monitor known metabolic patients on carnitine supplementation therapy |
CARN-P, CARN-S | Alberta Children's Hospital Biochemical Genetics | 14 days Test is performed on a weekly basis Contact for priority turn-around |
Liquid Chromatography, Electrospray-Ionization Tandem Mass Spectrometry (LC-MS/MS) with a deuterium-labelled internal standard | Determination of plasma or serum total and free carnitine levels, with calculated acylcarnitine fraction, enables the identification of a systemic carnitine deficiency (i.e., significantly low total and free carnitine levels with or without low acylcarnitine fraction, depending on underlying cause). Other alterations of total, free and/or acyl component measures of the test can be influenced by a patient’s underlying pathology, nutritional status as well as the application of carnitine supplementation therapy, and therefore such information should be provided on the requisition. Follow up testing of abnormal results may include recommendations for evaluation of the patient’s nutritional status, dietary predilections, current treatment and/or assessment of renal function This test may be reflexively performed on a sample in which a result from Acylcarnitine Profile (ACH) is suggestive of low free carnitine (a metabolite monitored but not reported by that test) Acylcarnitine Profile (ACH) testing may be reflexively performed based on evidence of unexplained increased acylcarnitine accumulation |
Blood | Collect 1 x 4.0 mL (0.25 mL plasma or serum min) sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label |
||||||||||||
| Further investigation of a patient with clear evidence of a systemic carnitine deficiency based on low plasma/serum total and free carnitine levels, to determine whether the cause of their deficiency is due to specific renal loss (i.e., carnitine uptake defect) or generalized renal insufficiency. Differentiating between primary and secondary cause can be evaluated by calculating fractional carnitine excretion | CARN-U | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a biweekly basis Contact for priority turn-around |
Liquid Chromatography, Electrospray-Ionization Tandem Mass Spectrometry (LC-MS/MS) with a deuterium-labelled internal standard | Urine | Collect 5 - 10 mL (1 mL min) random urine sample plain urine container Freeze |
If specimen is transferred to secondary aliquot tube or container, write "urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label |
||||||||||||||
| Cat-eye Syndrome 22q11.2 | COMT, HIRA, CECR2, IL17R | SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Real-Time PCR | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| CCND1 t(11;14) by FISH | CCND1 | Investigation of mantle cell lymphoma | University of Alberta Hospital Molecular Pathology | Test performed Monday to Friday | FISH | Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred. Alternatively, one slide stained with Hematoxylin & Eosin as well as six 4- micron baked (2 hours at 60°C) unstained charged slides can be submitted provide a copy of the corresponding pathology report |
Sodium heparin samples must be received within 48 hours of collection. On Fridays, samples must arrive prior to 1200H |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Molecular Pathology assesses sample integrity and pre-analytical processes for formalin-fixed, paraffin-embedded tissues |
|||||||||||||
| Blood and Bone Marrow | Collect 1 x 4 mL sodium heparin (green) tube - no plasma separator | |||||||||||||||||||||
| Charcot-Marie-Tooth Disease type 1A | PMP22 | CMT1A | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | MLPA: Designed to detect deletions and duplications, including the common 1.5-Mb duplication at 17p11.2 that accounts for 70-80% of individuals with Charcot-Marie-Tooth Disease type 1A Sanger sequencing: Analysis of all coding regions of PMP22 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - No restrictions Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics, Neurology |
|||||||||||||
| Charcot-Marie-Tooth Disease type 1B | MPZ | CMT1B | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | Sanger sequencing: Analysis of all coding regions of MPZ MLPA: Designed to detect deletions and duplications |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - No restrictions Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics, Neurology |
|||||||||||||
| Charcot-Marie-Tooth Disease X type 1 | Cx32 aka GJB1 | CMTX1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | "Sanger sequencing: Analysis of all coding regions of GJB1 MLPA: Designed to detect deletions and duplications" | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - No restrictions Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics, Neurology |
|||||||||||||
| Choroideremia | CHM | SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | Sanger Sequencing: Coding region of CHM gene MLPA: Detects partial or whole gene deletions and duplications |
Blood, DNA (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Chromosomal Microarray (CMA) | Prenatal primary indications for ultrasound anomaly(ies) and nuchal translucency greater than or equal to 3.5 mm Karyotype, amniotic fluid Cytogenetic testing, amniotic fluid Comparative genomic hybridization, amniotic fluid Array CGH, amniotic fluid |
CMAA (IN SUNQUEST), CGLNON (IN CERNER) | University of Alberta Hospital or Alberta Children's Hospital Cytogenetics | 2 weeks | Chromosomal Microarray | Amniotic Fluid | Collect minimum of 25.0 mL sterile polypropylene centrifuge tube | It is recommended that at the time of submission of the amniotic fluid, a blood sample on each parent should be collected and forwarded to the laboratory for possible parental studies All specimens should be received in the laboratory on the day of collection For UAH, specimens received on a Friday after 3:00pm will not get CMA on direct (uncultured specimen); CMA will be performed on cultured cells For ACH, specimens are received Monday through Friday: 8:00am to 4:15pm. Specimens received at the lab after 2:00pm will be processed the next working day |
MUST be submitted on a Chromosomal Microarray (CMA) Requisition form Ensure all sections of the requisition are complete Any amniotic fluid sample arriving with a request for only CMA testing, will have pre-analytical Rapid Aneuploidy Detection (RAD) testing added, and possibly reflex Karyotype testing if there is an abnormal RAD result, and any CVS sample will have pre-analytical RAD as well as Karyotype testing added Restricted to Obstetricians, Gynecologists or Geneticists For Possible Parental Studies, all requests MUST be submitted on a Molecular Diagnostic Laboratory Requisition form for CG North and for CG South, a Chromosomal Microarray (CMA) requisition form above |
|||||||||||||
| Prenatal primary indications for ultrasound anomaly(ies) and nuchal translucency greater than or equal to 3.5 mm Comparative Genomic Hybridization, chorionic villus sampling Array CGH, chorionic villus sampling |
CMACV (IN SUNQUEST), CGLNON (IN CERNER) | University of Alberta Hospital or Alberta Children's Hospital Cytogenetics | 3 weeks | Chromosomal Microarray | Chorionic Villus Sampling | Adequate amount of chorionic villi sampling to generate 15 mg minimum of typical villi after cleaning using sterile polypropylene centrifuge tube containing CVS transport media If request includes CMA and additional Molecular testing, 25 mg minimum of typical villi after cleaning |
||||||||||||||||
| Prenatal primary indications for ultrasound anomaly(ies) and nuchal translucency greater than or equal to 3.5 mm Fetal Tissue Array Comparative Genomic Hybridization, tissue (fetal) Array CGH, tissue |
CMATS (IN SUNQUEST) | University of Alberta Hospital Cytogenetics | 6 weeks | Chromosomal Microarray | Fetal Tissues | Collect minimum 1 cm3 preferred tissue specimen (umbilical cord, skin, spleen, lung, thymus, kidney or muscle) sterile 15 mL or 50 mL polypropylene centrifuge tube with NO fixative (DO NOT put tissue specimen in formalin) and refrigerate | Ship fresh tissue sample at 4 - 8oC within 48 hours of time of collection If unable to ship for receipt in lab in less than 48 hours from time of collection, freeze tissue sample with NO fixative and ship frozen specimen on dry ice Other samples are acceptable but quality of DNA obtained may not be optimal for this assay i.e., products of conception or placental tissue |
MUST be submitted on a Chromosomal Microarray (CMA) Requisition form Ensure all sections of the requisition are complete Restricted to Pediatricians, Pediatric Specialists, or Geneticists |
||||||||||||||
| Fetal Tissue Array | CGLNON (IN CERNER) | Alberta Children's Hospital Cytogenetics | 6 weeks | Chromosomal Microarray | Tissues | Collect minimum 1 cm3 preferred tissue specimen (umbilical cord, skin, spleen, lung, thymus, kidney or muscle) sterile 15 mL or 50 mL polypropylene centrifuge tube with NO fixative (DO NOT put tissue specimen in formalin) and refrigerate | For skin biopsy or surgical specimens, collect minimum sample required, DO NOT freeze, transport as quickly as possible If only placenta has been received by ACH Anatomical Pathology, a sample of the umbilical cord is preferred over a placenta sample Ship specimens at 4 - 8oC Placental or products of conception are at risk of maternal cell contamination and will be tested as a last resort if they are the only sample available. If the ONLY specimen sent are products of conception (POCs): Maternal cell contamination testing may be required to proceed with CMA testing, and may be requested once RAD testing is reported. If required, please submit a maternal blood sample (5mL EDTA and 5mL NaHep) with a Chromosomal Microarray (CMA) requisition form indicating 'for MCC studies'. Please also indicate the fetal patient information on the form |
MUST be submitted on a Chromosomal Microarray (CMA) Requisition form Ensure all sections of the requisition are complete Restricted to Pediatricians, Pediatric Specialists, or Geneticists for biopsies. Obstetricians, Pathologists and Geneticists for fetal tissues Multiple tissue specimens from a patient require a separate requisition for each specimen |
||||||||||||||
| Postnatal primary indication for isolated autism, isolated developmental delay, developmental delay and additional clinical features, congenital anomaly(ies) Comparative Genomic Hybridization, blood Array CGH, blood | CMAB (IN SUNQUEST), CGLMA (IN CERNER) | University of Alberta Hospital or Alberta Children's Hospital Cytogenetics | 4 weeks, Expedited 2 weeks | Chromosomal Microarray | Blood | For Patients >1 year of age - Collect 3.0 to 5.0 mL (1.0 mL min) in each of the required EDTA lavender) AND Sodium heparin tubes (dark green - NOT PST) For Patients |
Ship at room temperature. Specimens must be received by the Laboratory within 7 days of collection | MUST be submitted on a Chromosomal Microarray (CMA) Requisition form Ensure all sections of the requisition are complete Restricted to Pediatricians, Pediatric Specialists, or Geneticists For Products of Conception (POC’s) or Placental Tissues, a maternal blood sample (5ml EDTA) and a completed Molecular Diagnostic Laboratory Requisition (Edmonton) must also be submitted |
||||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Chromosome Analysis / Karyotype | Constitutional Cytogenetics Short-term DNA Storage |
CGLNON (IN CERNER) | Alberta Children's Hospital Cytogenetics | Routine 4 weeks, Stat 1 week Test set-up routinely Monday, Tuesday and Friday |
Blood | Collect 1 - 2 mL peripheral or cord blood for patient 6 months in a Sodium Heparin tube (dark green - NOT PST) | Must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete |
|||||||||||||||
| Constitutional Cytogenetics Short-term DNA Storage |
CGLNON (IN CERNER) | Alberta Children's Hospital Cytogenetics | 2 weeks Test set-up routinely Monday through Friday (excluding statutory holidays) |
Amniotic Fluid | 10 – 20 mL in sterile centrifuge tubes For possible microarray, minimum of 25 mL split into 3 centrifuge tubes |
Amniotic Fluid samples received for Karyotype testing will have the test cancelled and Rapid Aneuploidy Detection (RAD) testing will be performed instead If RAD testing is abnormal, karyotype testing will then be added on as a reflex test If a sample is received with indicating a family history of chromosomal abnormality or follow up chromosomal analysis, karyotype testing may be performed at the discretion of the Cytogeneticist |
Must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete |
|||||||||||||||
| 3 weeks Test set-up routinely Monday through Friday (excluding statutory holidays) |
Chorionic Villi | 10 – 30 mg in sterile container with transport media For possible microarray, minimum of 15 mg |
Chorionic Villi will also include RAD testing with karyotype testing | |||||||||||||||||||
| Constitutional Cytogenetics Cryopreservation of Cultured Cells |
CGLNON (IN CERNER) | Alberta Children's Hospital Cytogenetics | 4 weeks Test set-up routinely Monday through Friday (excluding statutory holidays) |
Tissues | Fibroblasts (skin biopsy, fresh surgical specimens) Sterile container containing transport media Alternate: Sterile container containing RPMI media |
DO NOT put tissue is formailn. DO NOT freeze specimen Fibroblasts (skin biopsy, fresh surgical specimens) For stillborn / POC fetal or placental tissues, refer to Rapid Aneuploidy Detection (RAD), fetal tissue Transport specimen at ambient temperature if within 1 hour of time of collection. If transport is delayed, transport at 4 - 8oC |
Must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete If a Chromosomal Microarray Analysis will also be requested on cultured tissue, also provide a completed Chromosomal Microarray (CMA) Requisition form Multiple specimens from a patient require separate requsition for each specimen |
|||||||||||||||
| Chromosome Breakage Studies | Fanconi Anemia | CGLBL (IN CERNER) | Alberta Children's Hospital Cytogenetics | 1 week Test set-up routinely Monday, Tuesday and Friday |
Blood | Collect peripheral or cord blood in sodium heparin (dark green - NOT PST) Newborn to 6 months: 1 - 2 mL 6 months to Adult: 3 - 4 mL |
Store whole blood specimen at room temperature | Must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete |
||||||||||||||
| CLL FISH Panel | Cancer cytogenetics | CBUN (CO PATH) | Alberta Children's Hospital Cytogenetics | 3 weeks Test set-up routinely Monday to Friday |
Blood | Collect 2.0 to 4.0 mL (1.0 mL min) blood in a sodium heparin tube (dark green - NOT PST) | Specimens should be received in the laboratory within 24 hours of time of collection Specimen MUST remain at room temperature |
Must be accompanied by a completed Hematology Specialty Requisition Ensure all relevant sections of the requisition are complete Specimens will be rejected if: -Primary tube received refrigerated or frozen -Any aliquot tube is received -Specimen is clotted or hemolyzed -Specimen is received in any tube other than sodium heparin -Insufficient specimen volume is received -Specimen received unlabelled -Specimen is received without a complete requisition |
||||||||||||||
| Combined Pituitary Hormone Deficiency 2 | PROP1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Sequence Analysis | Coding region of PROP1 | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Congenital Adrenal Hyperplasia | CYP21A2, CYP11B1 | CAH | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | MLPA: Detects dosage changes in CYP21A2 and CYP21P genes caused by deletions, duplications and gene conversions Sanger sequencing: Analysis of all coding regions of CYP21A2 and CYP11B1 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Congenital Central Hypoventilation Syndrome | PHOX2B | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Congenital Stationary Night Blindness | CACNA1F | CSNB | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.3166dupC (NM_005183) in the Mennonite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Costello Syndrome | HRAS | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Sanger sequencing | Sanger sequencing of the coding region of HRAS | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Craniosynostosis Panel | FGFR1, FGFR2, FGFR3, TWIST | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Targeted Sanger sequence analysis of exons with reported pathogenic variants in panel genes | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | "MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Perinatology and Pediatrics Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics" |
||||||||||||||
| CYLD Cutaneous Syndrome | CYLD | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||||
| Cystic Fibrosis | CFTR | Male Factor Infertility | SUNQUEST: CF-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 8 weeks | Targeted Mutation Analysis, Sequence and MLPA/Real Time | Targeted Mutation Analysis: 67 mutation CF kit + polyT Sequence and MLPA/Real Time: Full sequencing of coding region. Including detection of intron 8 TG tract for 5T carriers. MLPA/Real Time for detection of exon duplications and deletions MLPA/Real time only: Targeted deletion testing |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Full sequencing - Genetics, Respirology; Prenatal - Perinatal Clinic |
||||||||||||
| Cystine Screen | UCYST | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 14 days Test performed Monday to Friday |
Urine | Collect 5.0 ml (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
Positive tests will be confirmed by urine amino acid quantitation | Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
|||||||||||||||
| Cystinuria / Homocystinuria Screen | Initial investigation of disorders resulting in increased excretion of thiol-containing metabolites (i.e., homocystinuria and cystinuria). This test should not be used to monitor known homocystinuria or cytinuria patients on treatment as this is a qualitative test |
NP-U | Alberta Children's Hospital Biochemical Genetics | 8 days Test performed on a weekly basis Contact for priority turn-around |
Increased urinary homocystine or cystine excretion is detected colorimetrically with the addition of sodium cyanide and sodium nitroprusside. Positive and negative controls are run with patient samples | A positive nitroprusside test indicating the presence of either increased urine homocystine or cystine will result in reflexively tested amino acid and total homocysteine measurements on this sample, volume permitting | Urine | Collect 5 to 10 mL (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
If specimen is transferred to secondary aliquot tube or container, write "urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary container or aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Cystinuria Investigation | Diagnostic confirmation and monitoring of patients with cystinuria | UAACR | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 7 days Test performed Monday to Friday |
Urine | Collect 2.0 ml (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
Test requires creatinine result for interpretation | |||||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Deafness, Neurosensory; DFNB1 | GJB2 | Connexin 26 (FJB2-Related DFNB 1 Nonsyndromic Hearing Loss and Deafness) | SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Sequence Analysis | Sequencing of exon 2 | Blood, DNA, Prenatal and Fetal Tissue (see RAD) |
A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Deafness, Hereditary non-syndromic | GJB6 | Connexin 30 (GJB6-Related DFNB 1 Nonsyndromic Hearing Loss and Deafness) MTRNR1-Related Hearing Loss and Deafness Non-Syndrome Hearing Loss |
SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Real-Time PCR / Long range PCR | Detects common 309 kb and 232kb deletions | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Dentatorubral-Pallidoluysian Atrophy | ATN1 | DRPLA | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal, premutation and the majority of expanded pathogenic alleles; very large expanded pathogenic alleles will be detected by repeat-primed PCR but sizing will not be available | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| DICER1-Related Disorders | DICER1 | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||||
| Dilated Cardiomyopathy with Ataxia | DNAJC19 | 3-Methylglutaconic Aciduria Type 5 DCMA |
CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Sanger sequencing, including targeted sequence analysis | Sanger Sequencing: Analysis all of coding regions of gene DNAJC19 Targeted Sequence Analysis: Designed to detect c.130-1G>C (NM_145261) in the Hutterite population |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Duchenne Muscular Dystrophy | DMD | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology, Pediatrics and Cardiology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||||
| Dysequilibrium Syndrome | VLDLR | Cerebellar Ataxia, Mental Retardation and Dysequilibrium Syndrome | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Sanger sequencing and targeted deletion analysis by PCR | Sanger sequencing: Analysis all of coding regions of VLDLR PCR: Designed to detect the common 199kb deletion in the Hutterite population |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics and Neurology |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| EGFR Amplification by FISH | Investigation of gliomas | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Tuesday |
FISH | Slides are cut from paraffin block and are deparaffinized in xylene and dehydrated in ethanol. Specimen pretreatment is performed using SPoT-Light Tissue Pretreatment Kit (Life Technologies) and fluorescence in situ hybridization is performed using Vysis probes (Abbott Molecular Inc.) Vysis EGFR / CEP 7 FISH Probes hybridize to chromosome 7p12 and chromosome 7p11.1q11.1 (alpha satellites), therefore detecting increased copy number and amplification of the EGFR gene, as well as chromosome 7 polysomy. In normal cells, there is an equal number of red and green signals (ratio EGFR to CEP7 <1.2), but in abnormal cells a ratio of EGFR to CEP7 >=1.2 and >=10% of cells showing more than 3 EGFR signals is indicative of EGFR amplification All appropriate negative and positive controls are used. An average of 100 interphase nuclei were examined for each probe set by two observers using an Olympus Provis Fluorescence microscope system on 1000X magnification |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred. Alternatively, one slide stained with Hematoxylin & Eosin as well as six 4-micron baked (2 hours at 60°C) unstained charged slides can be submitted | Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Molecular Pathology assesses sample integrity and pre-analytical processes for formalin-fixed, paraffin-embedded tissues |
|||||||||||||
| EGFR Mutation Analysis | EGFR | Investigation of non-small cell lung cancers (NSCLC) | Real-time PCR | DNA extraction from FFPE is performed using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). The EGFR Mutation Assay is a Real-Time PCR assay performed using the QIAGEN EGFR RGQ PCR kit run on the Life Technologies QuantStudio instrument. This assay is capable of detecting Exon 19 deletions, L858R, L861Q, G719X (identifies but does not differentiate G719A/S/C), S768I, Exon 20 insertions, and T790M mutations The estimated sensitivity of this assay is 1-3% |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred. Alternatively, one slide stained with Hematoxylin & Eosin as well as five 10-micron non-baked unstained slides can be submitted | Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Molecular Pathology assesses sample integrity and pre-analytical processes for formalin-fixed, paraffin-embedded tissues |
||||||||||||||
| Endocrine Disorders Panel | AIP, CASR, CDC73, CDKN1B, MEN1, PRKAR1A, RET | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Ewing Sarcoma by FISH | EWSR1 | Investigation of Ewing Sarcoma EWSR1 gene rearrangement |
University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Monday and Wednesday (excluding statutory holidays) |
FISH | Slides are cut from paraffin block and are deparaffinized in xylene and dehydrated in ethanol. Specimen pretreatment and fluorescence in situ hybridization (FISH) is performed using Vysis® (Abbott Molecular Inc.) pretreatment kit and appropriate probes as described below The LSI EWSR1 dual colour probe consists of a 500 kb SpectrumOrange (red) labeled probe that flanks the 5’ side of the EWSR1 gene, and extends inward into intron 4. The second probe is a 1100 kb SpectrumGreen (green) probe, which flanks the 3’ side of the EWSR1 gene. The known breakpoints within the EWSR1 gene are restricted to introns 7 through 10 Normal nuclei contain two red/green (yellow) fusion signals (2F). In a normal cell that lacks a t(22q12) in the EWSR1 gene region, a two fusion signal pattern will be observed. In an abnormal cell with a simple t(22q12), the expected signal pattern will be one fusion, one red and one green signal This lab’s normal cut off is established at less than or equal to 10% of the cells showing an abnormal signal pattern. All appropriate negative and positive controls are used An average of 100 interphase nuclei are examined independently by two observers using an Olympus Provis Fluorescence microscope system on 1000X magnification |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred. Alternatively, one slide stained with Hematoxylin & Eosin as well as six 4-micron baked (2 hours at 60°C) unstained charged slides can be submitted | Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Molecular Pathology assesses sample integrity and pre-analytical processes for formalin-fixed, paraffin-embedded tissues |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Fabry Disease | Alpha-Galactosidase, leukocytes | FABRY | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed Monday to Friday |
Blood | Collect 1 x 6 mL or 3 mL (pediatric) Lithium Heparin (Light Green) or Sodium Heparin (Dark Green) – No Gel | Edmonton Zone Patients must be scheduled by 10am for blood collection at the Kaye Edmonton Clinic North/Central Zone Patients require a special arrangement including collection of a control sample and must phone the NMS/BG Lab (780-407-8820) in advance to discuss and scheduleMust receive samples before noon on the same day |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
||||||||||||||
| Factor V Leiden Mutation | Investigation of thrombophilia | APCGN | University of Alberta Hospital Molecular Pathology | 1 month Test performed on second Wednesday of each month |
Real-time PCR | After DNA isolation from peripheral blood sample, polymerase chain reaction is performed using the LightCycler 2.0 instrument and the Factor V Leiden kit purchased from F.Hoffman-La Roche Ltd. (Basel, Switzerland) The assay utilizes fluorogenic target-specific hybridization for genotyping of the amplified Factor V DNA and allows the detection of a single point mutation (G to A at position 1691) of the human Factor V gene. Data analysis is performed using the software purchased from the company |
Blood | Collect 1 x 4 mL EDTA (lavender) tube | Must be received in Molecular Pathology laboratory within 3 days of collection | Must be accompanied by a completed Hematology Specialty Requisition Please provide any pertinent clinical history on the requisition Factor V Leiden mutation requests must be ordered in conjunction with APC Resistance, Plasma Refer to Activated Protein C Resistance, plasma (APCE) for details of collection Submit request for both tests using the Hematology Specialty Requisition |
||||||||||||
| Factor XI deficiency | F11 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect p.E135X and p.F301L (NM_000128) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Familial Acute Myeloid Leukemia Panel | BEBPA, GATA2, RUNX1 | Cancer and Endocrine Next Generation Sequencing | SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Pathology | Up to 16 weeks | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on a NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. aCGH: Deletion / duplication testing is performed for those genes (except BEBPA) at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Familial Adenomatous Polyposis | APC | SUNQUEST: FAP-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 16 weeks | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on a NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. aCGH: Deletion / duplication testing is performed at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Familial Enteropathy with Villous Edema | APOA4 | SUNQUEST; FEVE-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Mutation Analysis | Detects c.552_749dup | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Familial Hypocalciuric Hypercalcemia | CASR | FHH | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Fanconi Anemia and DNA Repair Disorders Panel | ATM, BLM, BRCA2, BRIP1, FANCA, FANCB, FANCC, FANCD2, FANCE, FANCF, FANCG, FANCI, FANCL, FANCM, NBN, PALB2, RAD51C, REQL4, SLX4 | Cancer and Endocrine Next Generation Sequencing | SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 16 weeks | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on a NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. aCGH: Deletion / duplication testing is performed for those genes (except NBN) at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics, Hematology, Oncology, and Pediatrics |
||||||||||||
| Fascioscapulohumeral Muscular Dystrophy 1A | D4Z4 locus | FSHD | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 12-16 weeks | Southern Analysis | Detects deletion associated with FSHD. Haplotype analysis is NOT performed | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition ONLY available to Alberta residents Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Fecal Reducing Substances | Management of short bowel syndrome | FRED | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 5 days Test performed Monday to Friday |
Benedict’s Test | Does not detect sucrose Poor sensitivity for oligosaccharides False positive results can be seen with: Phenol, Salicylates, Chloral Hydrate, Aminoglycoside, Streptomycin, Isoniazid, Ascorbic Acid, Sulfonamide Antibiotics, Nalidixic Acid, Probenecid, Cephalosporins, Menthol, Penicillin |
Stool | Collect 5 mL (2 mL min) loose, random stool in sterile, screw top container Freeze immediately |
Please see Patient Collection Instructions: Fecal Reducing Substances and Fecal Sugar Identification | Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Rejection Criteria: Samples received more than 2 hours from collection not frozen must be cancelled by the collection site If pediatric lactase deficiency is suspected, order fecal sugar identification |
||||||||||||
| Fecal Sugar Identification | Pediatric lactase deficiency | SUGFE | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed Monday to Friday |
Does not detect sucrose | Stool | Collect 5 mL (2 mL min) loose, random stool in sterile, screw top container Freeze immediately |
Please see Patient Collection Instructions: Fecal Reducing Substances and Fecal Sugar Identification Ambient transport temperatures result in growth of bacteria which consume reducing sugars, this can yield a false negative result |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Rejection Criteria: Samples received more than 2 hours from collection not frozen must be cancelled by the collection site The breath hydrogen test is also available at the University of Alberta Hospital. If sucrase deficiency is suspected, order disaccharidase panel (requires small bowel biopsy sample) |
|||||||||||||
| Fibroblast Culture | UAH Test | University of Alberta Hospital Molecular Pathology | Set-up Monday through Friday (except for statutory holidays) | Biopsy | Preferrred - biopsy placed in a sterile culture medium (RPMI or 10% MEM) that is at AMBIENT temperature Alternative - biopsy can be placed in sterile saline. If the biospy has been placed in sterile saline, the biopsy needs to be delivered immediately to Molecular Pathology |
Allow medium to warm to ambient temperature for 30 minutes prior to use Biopsy must be received in Molecular Pathology by 1400H |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
|||||||||||||||
| ACH Test | FIBROBLAST | Alberta Children's Hospital Shared Genetic Laboratory Service | Set-up Monday through Friday (except for statutory holidays) | Biopsy | Biopsy specimen should be placed in transport media that is at AMBIENT temperature | Allow medium to warm to ambient temperature for 30 minutes prior to use Transport biopsy to Genetic Laboratory Services (ACH) during normal business hours (Monday through Friday, 8:00am to 4:00pm) Specimen can be refrigerated if not immediately being delivered for culture set-up |
Specimens can be submitted using either Biochemical Genetics Laboratory Requisition OR Constitutional Cytogenetics Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||||
| First Trimester Prenatal Screen | Includes pregnancy associated plasma protein A (PAPP-A) and free B hCG to estimate risk of trisomy 21 (Down sydrome), trisomy 18 (Edwards syndrome), and trisomy 13 (Patau syndrome) | FTPS | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed Monday to Friday |
Blood | Collect min 1 mL serum in plastic 3 mL No Gel (RED) clot activator Alternate: Plastic 5 mL SST Gel No Additive (GOLD) |
Spin and separate serum into plastic transport tube Store refrigerated. Transport at refrigerated temperature |
Edmonton Zone - Maternal Serum Screen Requisition MUST accompany specimen Please ensure that all requested information is completed FTPS is done at 11 weeks, 2 days - 13 weeks, 6 days gestation. Acceptable CRL range 45.0 - 80.9mm A nuchal translucency (NT) ultrasound is required and must be performed by an individual certified by the Fetal Medicine Foundation (FMF) to obtain NT measurements. The NT ultrasound is required on the same day, before the blood work is collected |
||||||||||||||
| FLT3 | FLT3 | Investigation of acute myelogenous leukemias (AML) | University of Alberta Hospital Molecular Pathology | 3 business days (5 calendar days) for new AML Test set-up twice a week |
PCR followed by capillary electrophoresis | After DNA extraction, PCR amplification is performed in two separate tubes for FLT3 and NPM1 using fluorescence labeled primers. Amplification products are subjected to fragment analysis by capillary electrophoresis using ABI 3130 genetic analyzer. Results are analyzed by Genemapper v3.2 software The sensitivity of these assays is 5% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Must be received in Molecular Pathology within 3 days of collection Keep samples refrigerated until ready for transport |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition CEBPA is a reflexed test that is only performed when the assay results for FLT3 / NPM1 are negative and the patient's karyotype is normal |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube |
Keep samples refrigerated until ready for transport | ||||||||||||||||||||
| Fluorescent In Situ Hybridization - FISH | Constitutional Cytogenetics | CGLBL (CERNER - FOR BLOOD) | Alberta Children's Hospital Cytogenetics | Routine 4 weeks, Stat 1 week Test set-up routinely Monday to Friday |
FISH | Blood (Peripheral or Cord) | Newborn to 6 months: 1 - 2 mL blood in a Sodium Heparin tube (dark green, not PST) 6 months to Adult: 3 - 4 mL blood in a Sodium Heparin tube (dark green, not PST) |
DO NOT centrifuge or separate specimen DO NOT refrigerate or freeze specimen Store and transport whole blood specimen at room temperature |
All specimens must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete, including indication of specific FISH probe/test Multiple specimens from a patient require a separate requsition for each specimen |
|||||||||||||
| CGLNON (CERNER - FOR ALL OTHER SAMPLE TYPES) | 2 weeks Test set-up routinely Monday to Friday |
Amniotic Fluid | 10 – 20 mL in sterile centrifuge tubes For possible microarray, minimum of 25 mL split into 3 centrifuge tubes |
DO NOT refrigerate or freeze specimen. Store and transport specimens at room temperature |
||||||||||||||||||
| 3 weeks Test set-up routinely Monday to Friday |
Chorionic Villi | 10 – 30 mg in sterile container with transport media For possible microarray, minimum of 15 mg |
||||||||||||||||||||
| 4 weeks Test set-up routinely Monday to Friday |
Skin / Tissue Biospy | 1 cm3 skin or tissue in a sterile container with tissue transport media | DO NOT put tissue in formailn DO NOT freeze Store and transport at 4 - 8oC Stillborn / POC fetal or placental tissue: refer to Rapid Aneuploidy Detection (RAD), fetal tissue (ACH) |
|||||||||||||||||||
| 1 week Test set-up routinely Monday to Friday |
Buccal Smear | 1 smear from each cheek on a separate glass slide. Buccal smears require a buccal smear collection kit | ||||||||||||||||||||
| Fragile X Syndrome | FMR1 | Fragile X - Associated Tremor/Ataxia Syndrome | SUNQUST: FRAX-; APPEND SPECIMEN TYPE CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE |
University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR; Methylation-sensitive Southern analysis | PCR analysis detects and sizes normal and premutation alleles, and detects but does not size expanded pathogenic alleles. Methylation-sensitive Southern analysis is performed when necessary to clarify pathogenicity of large premutations and small expanded alleles | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | For UAH, a maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
For MDL North (UAH), MUST be submitted on a Molecular Diagnostic Laboratory Requisition For MDL South (ACH), MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - No restrictions Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Friedreich Ataxia | FXN | FRDA | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal and premutation alleles only; detects the presence of expanded pathogenic alleles (no sizing available) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Frontal Temporal Dementia | C9ORF72 | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | PCR | Two PCR reactions to detect repeats in the 4 - 60 range | Blood, DNA (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Dx - Neurology Presymptomatic - Genetics |
||||||||||||||
| Fumarate Hydratase Deficiency | FH | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics and Metabolic specialist |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Galactocerebrosidase, T-lymphocyte | Investigation of Krabbe Disease (a.k.a., globoid cell leukodystrophy, galactosylceramidase deficiency, galactocerebrosidase deficiency) - a lysosomal storage disorder | GALC | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis Contact for priority turn-around |
Galactocerebrosidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | This procedure involves the establishment of an IL-2 stimulated T-lymphocyte culture and storage of cell pellets for the purposes of lysosomal enzyme testing | Blood | Collect 1 x 4.0 mL (3 mL plasma or serum min) sodium heparin tube (dark green) Alternate: Lithium heparin (dark green - NOT PST) |
Primary tube must remain at room temperature at all times It is recommended that patients go to ACH OP Lab for specimen collection The 3.0 mL of whole blood will be sufficient to test for multiple lysosomal enzymes in T-lymphocytes, if requested |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question Samples will be rejected if: -Primary tube not kept at room temperature -Any aliquot tube is received. -Any sample other than whole blood is submitted |
||||||||||||
| Galactokinase | Investigation of galactokinase deficiency | GK | Alberta Children's Hospital Biochemical Genetics | N/A Sample sent out on receipt |
Blood | Collect 1 x 4.0 mL EDTA tube (lavender) Alternate: -Sodium heparin (dark green - NOT PST) -Lithium heparin (dark green - NOT PST) -ACD A or ACD B (yellow) |
A separate specimen is required for this testing (i.e., cannot be combined with other test requests) This sample is stable for up to 10 days if refrigerated Keep sample refrigerated at all times following collection and during transport |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Condition suspected based on clinical presentation (e.g., congenital bilateral cataracts), or found to have excessive galactose in their urine by Sugar Chromatography (ACH). Classical galactosemia (i.e., galactose-1-phosphate uridyl transferase deficiency) should be ruled out by a normal Galactosemia Screen, Blood Spot (UAH) and/or Galactose-1-Phosphate Uridyltransferase (ACH) result prior to requesting this test or this test may be cancelled Samples will be rejected if: -Primary tube not refrigerated -Any aliquot is received. -Any sample other than whole blood is submitted |
||||||||||||||
| Galactose-1-Phosphate Uridyltransferase | Investigation of classic galactosemia (i.e., galactose-1-phosphate uridyltransferase deficiency) in a patient based on clinical presentation | G1PUT | Alberta Children's Hospital Biochemical Genetics | 14 days Test is performed on a biweekly basis Contact for priority turn-around |
Galactose-1-phosphate uridyltransferase activity is measured spectrophotometrically | Three enzymatic reactions are involved. In the first, endogenous galactose-1-phosphate uridyltransferase (G1PUT) in the RBC lysate catalyzes the reaction between galactose-1-phosphate (G1P) and Uridine-5’-diphosphoglucose (UDPG) to produce glucose-1-phoshate. Two additional reactions are performed in which an increase in absorption of NADPH at 340nm is directly proportional to the glucose-1-phosphate derived from the catalytic activity of G1PUT in the sample | Blood | Collect 1 x 4.0 mL (3 mL plasma or serum min) sodium heparin tube (dark green) Alternate: Lithium heparin (dark green - NOT PST) |
It is recommended that patients go to ACH OP Lab for specimen collection | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Not refrigerated -Not received in primary collection tube -Any sample other than heparinized whole blood is submitted For Alberta-born patients: Samples received for Galactose-1-Phosphate Uridyltransferase testing will be further investigated to determine if a newborn screen result is available For Alberta-born patients with no available newborn screen result and for non-Alberta-born patients a bloodspot card will be sent to the University of Alberta Newborn Metabolic Screen & Biochemical Genetics Lab for a Galactosemia Screen For monitoring known galactosemic patients on dietary therapy, please order Galactose-1-Phosphate, Quantitative. |
||||||||||||
| Galactosemia | GALT | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.563A>G (p.Q188R) (NM_000155) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Galactosemia Screen, blood spot | Initial investigation of galactosemia | GALBS | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed Monday to Friday |
Blood spot | Newborn Metabolic Screening Blood Spot Requisition (card) | Ensure that expiry date of Newborn Metabolic Screening card (requisition) has not been exceeded before proceeding with collection Collect two saturated blood spots on the Newborn Metabolic Screening card Use a heel poke if patient is less than one year old or a finger poke if older Could also collect in a Lithium heparin or Sodium heparin tube at any age and apply whole blood to newborn screening card using a capillary tube. Do not allow the capillary tube to contact the card. Each blood spot should require less than 100uL of blood Allow the newborn metabolic screening card to air dry for a minimum of 3 hours Write "ACYLCARNITINE, PROFILE" on the newborn metabolic screening card Ship at ambient temperature in a biohazard bag |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Rejection cirteria: Blood spots for Galactosemia screen are stable for up to one month only Please see the Lab Bulletin for Galactosemia testing in Alberta For confirmation of galactosemia in a patient, please order Galactose-1-Phosphate Uridyl Transferase. For monitoring of known galactosemia patients on dietary therapy, please order Galactose-1-Phosphate, quantitation Prior blood transfusion may give a falsely normal result |
||||||||||||||
| Gastrointestinal / Polyposis Panel | APC, BAP1, BMPR1A, CDH1, CHEK2, EPCAM, MLH1, MSH2, MSH6, MUTYH, PMS2, POLD1, POLE, PTEN, RHBDF2, SMAD4, STK11, TP53 | Cancer and Endocrine Next Generation Sequencing | SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Pathology | Up to 16 weeks | NGS and Exon Level Array CGH For PMS2 (exons 11-15 only) Sanger Sequencing and MLPA |
NGS: Genomic DNA is sequenced on a NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS aCGH: Deletion / duplication testing is performed for all genes (except PMS2, POLD1, POLE, RHBDF2) at single exon-level resolution MLPA: Detects deletions and duplication Sanger Sequencing: Analysis of PMS2 exons 11-15 only |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
||||||||||||
| Gastrointestinal Stromal Tumor Panel | KIT, PRKAR1A, SDHA, SDHB, SDHC, SDHD | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Gaucher Disease | Beta-Glucocerebrosidase Beta-Glucosidase, leukocytes |
GAUCH | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics |
4 days Test performed Monday to Friday |
Blood | Collect 1 x 6 mL or 3 mL (pediatric) Lithium Heparin (Light Green) or Sodium Heparin (Dark Green) – No Gel | Edmonton Zone Patients must be scheduled by 10am for blood collection at the Kaye Edmonton Clinic North/Central Zone Patients require a special arrangement including collection of a control sample and must phone the NMS/BG Lab (780-407-8820) in advance to discuss and scheduleMust receive samples before noon on the same day |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
||||||||||||||
| Glaucoma | CYP1B1, MYOC | Congenital Glaucoma | SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis /td> | Analysis of exon 1-3 of MYOC and Exons 2-3 of CYP1B1 | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Glycosaminoglycan (Mucopolysaccaride) Screen | Initial investigation for mucopolysaccharidoses (a.k.a., disorders of glycosaminoglycan catabolism) | MPSSCREEN | Alberta Children's Hospital Biochemical Genetics | 8 days Test is performed on a weekly basis Contact for priority turn-around |
Increased urinary mucopolysaccharides are detected using both toluidine blue spot and cetyltrimethylammonium-bromide (CTMAB) turbidity tests | Positive toluidine blue and CTMAB tests indicating the presence of increased urine mucopolysaccharides (a.k.a., glycosaminoglycans or GAGs) will result in reflexive quantitative uronic acids and GAG electrophoresis testing on the same sample, volume permitting. Positive and negative controls are run with patient samples | Urine | Collect 5 - 10 mL (4.0 mL min) of random urine sample in plain urine container Freeze immediately |
Prefer first morning void If specimen is transferred to secondary aliquot tube or container, write ""urine"" on label |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test should not be used to monitor known mucopolysaccharidosis patients on treatment as this is a qualitative test Samples will be rejected if: -Primary container or aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Glycosaminoglycan Electrophoresis, Qualitative | Suspicion of a mucopolysaccharidosis in a patient (in conjunction with quantitative uronic acids), or when the initial glycosaminoglycan (mucopolysaccharide) screen has been found to be abnormal and there was insufficient sample for reflexive testing | GAG | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis Contact for priority turn-around |
Discontinuous one-dimensional cellulose acetate electrophoresis | The presence of abnormal glycosaminoglycans dermatan and/or heparan sufate, or keratan sulfate, in addition to the normal urine constituents chondroitin 4 and 6 sulfates, is highly suggestive of a biochemical diagnosis of a mucopolysaccharidosis. Such a finding should be supported by an elevated quantitative uronic acid level in the majority of cases. Further enzyme or molecular-based testing is required to confirm the specific enzyme deficiency in each case | Urine | Collect 5 - 10 mL (4.0 mL min) of random urine sample in plain urine container Freeze immediately |
Prefer first morning void If specimen is transferred to secondary aliquot tube or container, write ""urine"" on label |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided. This test is for the investigation of mucopolysaccharidoses and is required to be performed in conjunction with Uronic Acids, Quantitative (ACH). Minimum volume is for both tests Samples will be rejected if: -Primary container or aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Half Cystine Quantitation | Investigation of cystinosis, and for monitoring known cystinosis patients on cysteamine (e.g., Cystagon, Procysbi) therapy | CYSTINOSIS | Alberta Children's Hospital Biochemical Genetics | 14 days Prior arrangement is required before ordering Contact for priority turn-around |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with a deuterium-labelled internal standard | Proper interpretation of results in known cystinosis patients on treatment is dependent on the timing of the patient’s last dose | Blood | Collect 2 x 4.0 mL sodium heparin tube (dark green - NOT PST) | It is recommended that patients go to ACH OP Lab for specimen collection. Tubes must be FULL The ACH Biochemical Genetics Lab (403-955-7379) must be notified at least 24 hours before sample collection to ensure a member of the lab is present to immediately process the specimen |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided |
||||||||||||
| HBA-related disorders | HBA1, HBA2 | Alpha thalassemia Constant Spring HBA1 HBA2 Hemoglobinopathy Hydrops fetalis Hb Barts |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Multiplex PCR and Sanger sequencing | Multiplex PCR: Detects - alpha3.7 (rightward), - alpha4.2 (leftward), --SEA, --FIL, --MED, -(alpha)20.5, --THAI Sanger sequencing: Analysis of all coding regions of HBA1 and HBA2 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Hemoglobinopathy screen results are required |
|||||||||||||
| HBB-related disorders | HBB | Hemoglobinopathy HbS Beta thalassemia Sickle cell anemia HbC HbE |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | Sanger sequencing: Sequence analysis of the entire globin gene (introns and exons), the promoter region and the CAP regions & common 619 bp deletion. Detects HbC, HbD, Hb E, HbS, etc. MLPA: Detects deletions and duplications |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Hemoglobinopathy screen results are required |
|||||||||||||
| Hemophilia A | F8 | Factor VIII Deficiency | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Inverse Shifting PCR | Detects intron 22 inversions and deletions and intron 1 inversion | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Hemoglobinopathy screen results are required |
|||||||||||||
| Hereditary Diffuse Gastric Cancer | CDH1 | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 16 weeks | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on an NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS aCGH: Deletion/duplication testing is performed at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Hereditary Fructose Intolerance | ALDOB | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect the following common pathogenic variants (NM_000035): c.448G>C (p.A150P) c.524C>A (p.A175D) c.1005C>G (p.N335K) |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Hereditary Hemochromatosis | HFE | HFE HH |
SUNQUEST: HC-; APPEND SPECIMEN TYPE CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE |
University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | 2 weeks | Targeted Sequence Analysis | Detects c.845G>A (p.Cys282Tyr) and c.187C>G (p.His63Asp) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception For UAH, Chorionic Villi specimens cannot be received on Fridays |
For MDL North (UAH), MUST be submitted on a Molecular Diagnostic Laboratory Requisition For MDL South (ACH), MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical |
||||||||||||
| Hereditary Multiple Osteochrondromatosis Panel | EXT1, EXT2 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Hereditary Neuropathy with liability to Pressure Palsies | PMP22 | HNPP | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | MLPA: Detects deletions and duplications Sanger sequencing: Analysis of all coding regions of PMP22 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | "MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - No restrictions Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Neurology and Genetics" |
||||||||||||
| Hereditary Multiple Osteochrondromatosis Panel | EXT1, EXT2 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Hereditary Neuropathy with liability to Pressure Palsies | PMP22 | HNPP | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | MLPA: Detects deletions and duplications Sanger sequencing: Analysis of all coding regions of PMP22 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - No restrictions Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Neurology and Genetics |
||||||||||||
| Hexosaminidase | Diagnostic investigation of Tay-Sachs or Sandhoff disease Tay-Sachs disease |
HEX-S | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Hexosaminidase activity is measured fluorometrically using a 4-methylumbelliferyl-linked substrate | Blood | Collect 1 x 4.0 mL (0.5 mL serum min) in non-additive (red top - NOT SST) Centrifuge, transfer serum to aliquot tube, write ""Red Top"" on the aliquot tube label, and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Aliquot received thawed -Tube type not written on aliquot tube label -Plasma or urine sample submitted For preconception testing for Tay-Sachs and Sandhoff disease carrier status, see Tay-Sachs CARRIER Testing (ACH) |
||||||||||||||
| Hidrotic Ectodermal Dysplasia 2 | GJB6 | Clouston Syndrome GJB6 |
SUNQUEST: MDG-; APPEND TET NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Sequencing | Sequencing of exon 3 | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Hirschsprung disease | RET | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Homocysteine | Investigation of a primary disorder associated with methionine or cobalamin metabolism based on clinical presentation, or when an initial cystinuria/homocystinuria (a.k.a., nitroprusside) screen was found to be abnormal and there was insufficient sample for reflexive testing Total homocysteine can be elevated in the following enzyme deficiencies: Disorders Detectable by Elevated Plasma Total Homocysteine: -cystathionine beta-synthase deficiency (classical homocystinuria) -methylenetetra-hydrofolate reductase deficiency (MTHFR) -methionine synthase deficiency -Cobalamin (Cbl) C, D, E, F, G, J and X deficiencies -transcobalamin II deficiency -S-adenosylhomocysteine hydrolase (SAHH) deficiency -glycine N-methyltransferase (GNMT) deficiency -methionine adenosyltransferase (MAT) I/III deficiency -adenosine kinase (ADK) deficiency Other causes of Elevated Plasma Total Homocysteine: -nutritional vitamin B12 deficiency -folate deficiency -nutritional B6 deficiency |
THCYS-P, THCYS-S | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis Contact for priority turn-around |
Reduction using liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with a deuterium-labelled internal standard | Blood | Collect 1 x 4.0 mL (0.25 mL plasma min) sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided. Blood test only available for pediatric patients (less than 16 years of age) and monitoring of known homocystinuria patients; all other types of testing should refer to the Calgary Lab Services Guide to Lab Services entry Homocysteine Samples will be rejected if: -Primary tube not refrigerated (blood) -Aliquot tube received thawed -Tube type not written on aliquot tube label This test should not be ordered for the investigation of nutritional vitamin B12 deficiency, nor for assessing carrier status of the common methylene tetrahydrofolate reductase (MTHFR) polymorphisms |
|||||||||||||
| THCYS-U | Urine | Collect 5 - 10 mL (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
Prefer first morning void If specimen is transferred to secondary aliquot tube or container, write ""urine"" on label |
|||||||||||||||||||
| Huntington Disease | HTT | Huntington Chorea HD |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal and premutation alleles only; detects the presence of expanded pathogenic alleles (no sizing available) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology and Psychiatry Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Hydroxybutyrate, beta | Suspected of having an inborn error of metabolism affecting energy production (e.g., hepatic glycogen storage disease) or utilization (e.g., fatty acid oxidation disorders). This test can also be used to monitor adherence of patients treated with the ketogenic diet. This test may also help in the evaluation of diabetes, as well as other conditions leading to or resulting from carbohydrate deprivation, reduced carbohydrate utilization, ketoacidosis, alkalosis or altered cellular redox state Ketosis is a common feature in acutely ill patients and in some subjects can result in severe life threatening metabolic acidosis |
3HB | Alberta Children's Hospital Biochemical Genetics | 7 days Test is performed on a weekly basis Contact for priority turn-around |
Beta-hydroxybutyrate is detected spectrophotometrically using a commercially available kit | Metabolite reported: D-3-hydroxybutyrate | Blood | Collect 1 x 4.0 mL (0.25 mL plasma min) sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Sodium fluoride (grey) |
Collection after an 8 to 15 hour fast preferred (but not mandatory) Indicate length of fast on the requisition (with collection time) If patient is on ketogenic diet, indicate whether sample is fed or fasted Physician test requests from known metabolic patients (non-fasting) are also acceptable At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label |
||||||||||||
| Hypokalemic Periodic Paralysis | CACNA1S, SCN4A | SUNQUEST: HOKPP-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Sequence Analysis | Sequence exons 11 and 30 / sequence exon 12 | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Hypoxanthine Guanine Phosphoribosyltransferase | Further investigation of a patient with a high clinical suspicion, or initial laboratory results consistent with, Lesh-Nyhan syndrome (a.k.a, hypoxanthine guanine phosphoribosyltransferase deficiency) | HPRT | Alberta Children's Hospital Biochemical Genetics | N/A Sample sent out on receipt |
Blood Spot | 2 full blood spots | Self-collection of a sample by a patient using the pre-labelled blood spot card requires previous instruction by an ACH metabolic physician or an ACH metabolic dietician If a patient brings a pre-labelled blood spot card to the collection lab, warm a finger or heel before collection and fill a minimum of two circles on the card using a single drop of whole blood (blood must soak through to the back of the card Do NOT smear blood onto the card or apply to both sides of card. If a single drop is insufficient, a second drop can be applied Do NOT use a capillary tube to transfer blood to the card Allow card to dry for a minimum of 4 hours after collection by clipping or lean card against an object on a clean, flat, non-absorbent surface. Keep card away from sunlight and heat. Place dry collection card inside a biohazard bag Transport at room temperature |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided This test should NOT be used as a primary screen and may be cancelled if not adequately justified on the requisition. Confirmation of hyperuricemia (see Urate - Urine, CLS) and/or evidence of elevated urine hypoxanthine, xanthine and uric acid (see Purine and Pyrimidine Profile - ACH) are required for testing to proceed |
||||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| IgH Gene Rearrangement | IgH | Investigation of lymphoproliferative disorders B Cell Gene Rearrangement |
University of Alberta Hospital Molecular Pathology | 1 week Test set-up on Monday and Thursday (excluding statutory holidays) |
PCR followed by capillary electrophoresis | DNA is extracted using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). A commercially available kit is purchased from InVivoScribe Technologies (San Diego, California) and utilized in this assay. This assay has been validated internally using >30 B-cell lymphomas or benign lymphoid tissues for which the diagnoses were confirmed histologically and immunophenotypically PCR is performed using Invitrogen Platinum High Fidelity Taq with fluorescence-labelled primers in three different mixes. The PCR products are subjected to capillary electrophoresis on an Applied Biosystems 3130/3500XL Genetic Analyzer using fragment analysis GeneMapper software. The integrity of the DNA is assessed using a Specimen Control Size Ladder detecting four different fragment sizes. Appropriate polyclonal and monoclonal controls (B-cell lymphoma cell lines) are used. A no template control is also included This assay does not detect 100% clonal cell populations and cannot detect less than 5 positive cells/95 normal cells The clinical sensitivity of this assay is determined to be 75% and clinical specificity is 100% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Whole blood samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Test requests are restricted to hematopathologists and lymphoma pathologists |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | |||||||||||||||||||||
| Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | ||||||||||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| JAK2 Exon 12 Mutation | JAK2 | Investigation of myeloproliferative neoplasms | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Wednesday |
PCR and sequence analysis | DNA is extracted using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). PCR amplification is performed using M13 tailed primers. Bidirectional sequencing of the product is performed using the BigDye Direct Cycle Sequencing kit (Life Technologies) and run on the Applied Biosystems 3500xL Genetic Analyzer. The obtained DNA sequence is then analyzed using ABI SeqScanner® and SeqPilot® software programs and compared to the published sequences in the National Center for Biotechnology Information (NCBI) database at www.ncbi.nlm.nih.gov. (GenBank) Analytical sensitivity is 20% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Samples must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| JAK2 V617F Mutational Analysis | JAK2 | Investigation of myeloproliferative neoplasms, namely polycythemia vera, essential thrombocythemia and primary myelofibrosis | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Monday (excluding statutory holidays) |
Real-time PCR | After DNA extraction, the qualitative JAK2 V617F assay is performed using the JAK2 MutaScreenTM Kit (Ipsogen Inc., New Haven, CT). The JAK2 gene sequence spanning the mutation site was amplified using real time polymerase chain reaction with two allele-specific probes that allow detection of the presence of the JAK2 V617F mutation The sensitivity of the assay is 2% (2% JAK2 V617F DNA in a normal background) |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition The mutation may be undetectable in samples containing low numbers of granulocytes or monocytes, or in the presence of a high lymphocyte population, as lymphoid cells generally do not harbor the acquired JAK2 V617F mutation |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Samples must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| Joubert Syndrome and Related Disorders | TMEM237 | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.52C>T (NM_001044385) in the Hutterite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology and Psychiatry Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Juvenile Hemochromatosis | HFE2A | HJV Hemochromatosis Type 2 |
University of Alberta Hospital Molecular Diagnostics | Real-Time PCR | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Contact laboratory |
||||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Karyotype | Chromosome analysis of blood Cytogenetic testing of blood Turner Syndrome |
CGB (CO PATH) | University of Alberta Hospital Cytogenetics | 4 weeks Test set-up routinely Monday to Friday |
Blood | Collect Sodium Heparin tube (dark green - NOT PST) Newborn to 6 months: 2 mL peripheral blood Children: 3 mL (2.0 mL min) peripheral blood Adult: 4 mL (2.0 mL min) peripheral blood |
Must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete Specimens will be rejected if: -Primary tube received refrigerated or frozen -Any aliquot tube is received -Specimen is clotted or hemolyzed -Specimen is received in any tube other than sodium heparin -Insufficient specimen volume is received -Specimen received unlabelled -Specimen is received without a complete requisition |
|||||||||||||||
| Chromosome analysis of unstimulated peripheral blood Cytogenetic testing of unstimulated peripheral blood |
CBUN (CO PATH) | 3 weeks Test set-up routinely Monday to Friday |
Unstimulated Peripheral Blood | Collect a 2.0 - 4.0 mL (1.0 mL min) of blood Sin sodium heparin (dark green - NOT PST) | For unstimulated peripheral blood specimens on urgent cases collected on weekends or statutory holidays page the Clinical Genetic Technologist on call at 780-445-6036 | Must be accompanied by a completed Hematology Specialty Requisition Ensure all relevant sections of the requisition are complete If multiple myeloma is suspected, this must be indicated on the requisiton Order Restrictions: Specimens must have dividing cells Specimens will be rejected if: -Primary tube received refrigerated or frozen -Any aliquot tube is received -Specimen is clotted or hemolyzed -Sample is received in any tube other than sodium heparin -Insufficient sample volume is received. -Specimen received unlabelled -Specimen is received without a complete requisition |
||||||||||||||||
| Chromosome analysis of bone marrow Cytogenetic testing of bone marrow |
CGBM (CO PATH) | 3 weeks Test set-up routinely Monday to Friday |
Bone Marrow | Collect 1.0 - 3.0 mL of bone marrow into 15 mL centrifuge tube with transport media Alternate: Sodium heparin (dark green - NOT PST) |
Preferrably, specimens should be received in the laboratory on the day of collection Specimens will be accepted up to 24 hours after time of collection For bone marrows on urgent cases collected on weekends or statutory holidays page the Clinical Genetic Technologist on call at 780-445-6036 |
Must be accompanied by a completed Hematology Specialty Requisition Ensure all relevant sections of the requisition are complete Specimens will be rejected if: -Primary tube received refrigerated or frozen -Any aliquot tube is received -Sample is received in any tube other than transport media or sodium heparin. -Insufficient sample volume is received -Specimen received unlabelled -Specimen is received without a complete requisition |
||||||||||||||||
| Chromosome analysis of chorionic villi Cytogenetic testing of chorionic villi |
CGCV (CO PATH) | 3 weeks Test set-up routinely Monday to Friday |
Chorionic Villus Sampling | 15 mg minimum of typical villi after cleaning for RAD and chromosome analysis or 25 mg minimum of typical villi after cleaning for RAD, chromosome analysis and Molecular testing in sterile polypropylene centrifuge tube containing CVS transport media | Must be accompanied by a completed Constitutional Cytogenetics Requisition Ensure all relevant sections of the requisition are complete Specimens will be rejected if: -Primary tube received refrigerated or frozen -Specimen is received in any tube other than CVS transport media -Insufficient specimen volume is received -Specimen received unlabelled -Specimen is received without a complete requisition |
|||||||||||||||||
| KIT for AML | KIT | Investigation of acute myeloid leukemia | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Wednesday |
PCR and sequence analysis | DNA is extracted using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). PCR amplification is performed using M13 tailed primers. Bidirectional sequencing of the product is performed using the BigDye Direct Cycle Sequencing kit (Life Technologies) and run on the Applied Biosystems 3500xL Genetic Analyzer. The obtained DNA sequence is then analyzed using ABI SeqScanner® and SeqPilot® software programs and compared to the published sequences in the National Center for Biotechnology Information (NCBI) database at www.ncbi.nlm.nih.gov. (GenBank) Analytical sensitivity is 20% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Samples must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| KIT for Mastocytosis | KIT | Investigation of systemic mastocytosis | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Wednesday |
PCR and capillary electrophoresis | DNA is extracted on the QIAsymphony instrument. Allele specific PCR amplification is performed in two separate tubes for mutant and wildtype KIT using fluorescently labeled primers. Amplification products are subjected to fragment analysis by capillary electrophoresis using ABI 3130/3500 genetic analyzer. Results are analyzed by Genemapper v3.2 software The sensitivity of this assay is 1% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Samples must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | ||||||||||||||||||||
| KIT for Melanoma / GIST | KIT | Investigation of melanoma When ordered in conjunction with PDGFRA used for the investigation of Gastrointestinal Stromal Tumours (GIST) |
University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Wednesday |
PCR and sequence analysis | DNA is extracted using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). PCR amplification is performed using M13 tailed primers. Bidirectional sequencing of the product is performed using the BigDye Direct Cycle Sequencing kit (Life Technologies) and run on the Applied Biosystems 3500xL Genetic Analyzer. The obtained DNA sequence is then analyzed using ABI SeqScanner® and SeqPilot® software programs and compared to the published sequences in the National Center for Biotechnology Information (NCBI) database at www.ncbi.nlm.nih.gov. (GenBank) Analytical sensitivity is 20% |
Paraffin-Embedded Tissue | "A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted" | Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Krabbe Disease | GALC | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted deletion analysis by PCR | Designed to detect the common 30kb deletion in the European population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| KRAS Mutation Analysis | KRAS | Investigation of colon cancer | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Monday and Wednesday (excluding statutory holidays) |
PCR followed by capillary electrophoresis | DNA extraction from FFPE samples is performed using the QIAGEN QIAsymphony DSP DNA mini kit. Analysis for BRAF, KRAS, NRAS mutations is performed after PCR amplification using a single base extension technique (SNaPshot multiplex kit by Life Technologies) and capillary electrophoresis on the 3130 Genetic Analyzer. This multiplexed assay is capable of detecting mutations at BRAF codon 600 and KRAS/NRAS codons 12, 13, 61, 117, and 146 The estimated sensitivity of this assay is 10% |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Leigh Syndrome | NDUFS4 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.393dupA (NM_002495) in the Hutterite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Li-Fraumeni Syndrome | TP53 | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE |
University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception For UAH, Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Limb-girdle Muscular Dystrophy Type 2H | TRIM32 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.1459G>A (p.D487N) (NM_012210) in the Hutterite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) |
A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Limb-girdle Muscular Dystrophy Type 2I | FKRP | Muscular Dystrophy - Dystroglycanopathy Type C | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.826C>A (p.L276I) (NM_024301) in the Hutterite and European populations | Blood, DNA, Prenatal and Fetal Tissue (see RAD) |
A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Long Chain 3-Hydroxyacyl-CoA Dehydrogenase Deficiency / HELLP | HADHA | SUNQUEST: LCAD-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Mutation Analysis | Detects the c.1528G>C mutation | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Lynch Syndrome Cancer Panel | MLH1, MSH2, MSH6, PMS2, EPCAM | Cancer and Endocrine Next Generation Sequencing | SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Pathology | Up to 16 weeks | NGS and Exon Level Array CGH For PMS2 - Sanger Sequencing and MLPA |
MLH1, MSH2, MSH6, PMS2 (exons 1-10 only) by NGS: Genomic DNA is sequenced on a NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS EPCAM (exons 3. 8, and 9 only), MLH1, MSH2, MSH6 by aCGH: Deletion / duplication testing is performed for those genes indicated at single exon-level resolution PMS2 by MLPA: Detects deletions and duplications PMS2 by Sanger Sequencing: Sequence analysis if PMS2 exons 11-15 only |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Maturity Onset Diabetes of the Young (MODY) Type 3 | HNF1A | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.1137delT (NM_000545) in the Hutterite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Medium Chain Acyl-Coenzyme A Dehydrogenase Deficiency | MCAD | SUNQUEST: LCAD-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Mutation Analysis | Detects c.985A>G (Lys304Glu) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Methylmalonic Acid | Plasma methylmalonic acid can aid in the diagnosis of functional B12 deficiency when the plasma B12 levels are in the low-normal range (150 to 400 pmol/L) Urine methylmalonic acid can aid in differentiating between a true functional B12 deficiency and renal insufficiency, both of which can lead to a mild to moderate -elevation of plasma methylmalonic acid Methylmalonic acid can be elevated in the following enzyme deficiencies / disorders: -methylmalonyl-CoA mutase deficiency -methylmalonyl-CoA epimerase deficiency -cobalamin (Cbl) A, B, C, D, F, J and X deficiencies -intrinsic factor deficiency and Imerslund-Grasbeck syndrome -transcobalamin II deficiency -ACSF3 deficiency (a.k.a., combined malonic and methylmalonic aciduria or CMAMMA) -SUCLA2 and SUCLG1 deficiencies Other causes of elevated methylmalonic acid: -nutritional vitamin B12 deficiency -renal insufficiency (plasma/serum only) |
MMA-P, MMA-S, MMA-U | Alberta Children's Hospital Biochemical Genetics | 14 days Test is performed on a weekly basis Contact for priority turn-around |
Blood | Collect 1 x 4.0 mL sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) -Random specimen: plain urine container |
Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Primary tube not refrigerated -Primary urine container not refrigerated or frozen -Aliquot tube received thawed -Tube type not written on aliquot tube label Plasma methylmalonic acid test should only be ordered when there is a high index of clinical suspicion of a nutritional B12 deficiency, based on patient presentation Urine methylmalonic acid test should not be ordered in conjunction with urine organic acids, as this metabolite is also reported as part of that test, and this test will be reflexively performed on any urine sample with elevated levels detected |
||||||||||||||
| Urine | Collect 5 - 10 mL (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
If specimen is transferred to secondary aliquot tube or container, write "urine" on label | ||||||||||||||||||||
| Microsatellite Instability | Investigation of colonic, ovarian and uterine adenocarcinomas | University of Alberta Hospital Molecular Pathology | 1 month Test performed every second week |
PCR followed by capillary electrophoresis | The assay involves multiplex PCR of five microsatellite (MS) loci for both normal and tumor tissue. The MS loci tested include mononucleotide repeats BAT25, BAT26, BAT40, CAT25 and NR21 GUIDE TO TEST RESULT: Comparative evaluation of normal and tumor peak patterns may reveal shifts in one or more MS sequences amplified in the tumor, representing microsatellite instability (MSI). Tumor microsatellite instability is classified as follows: high frequency instability (MSI-H) when two or more sequences demonstrate instability, low frequency instability (MSI-L) if one of the sequences demonstrate instability, and microsatellite stable (MSS) when no shifts are evident |
Paraffin-Embedded Tissue | Representative formalin-fixed, paraffin-embedded tissue block(s) of tumor and normal (uninvolved) tissue is preferred. A 1 x 4 mL EDTA blood sample may also be used as a normal specimen Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Test requires comparison between tumor and normal specimens. Test will not be performed if a normal tissue or blood sample is not submitted |
|||||||||||||
| Mitochondrial Disease | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Mutation Analysis, Southern and Sequence Analysis | Targeted Mutation Analysis: Detects mutations associated with MERFF, MELAS, NARP Targeted Mutation Analysis: Detects the following mutations associated with Lebers Hereditary Optic Neuropathy 3 mutations- m.3460G>A , m.11778G>A, m.14484T>C Southern: Detects deletion syndromes Sequence Analysis: Full screen |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Dx - Metabolic and Neuromuscular specialists (for deletion syndromes and full screen) |
||||||||||||||
| MLH1 Promoter Hypermethylation | MLH1 | Distinguishes sporadic MSI-High tumors from tumors with germline mutations in the MLH1 mismatch repair gene (Lynch Syndrome) | University of Alberta Hospital Molecular Pathology | 1 month Test performed every three weeks |
Bisulfite modification, PCR followed by capillary electrophoresis | PCR-based assays for the detection of hypermethylation of the MLH1 promoter region and BRAF V600E mutation. The methylation specific PCR assay is a modification of the method described by Grady et al. (Grady WM, Rajput A, Lutterbaugh JD, Markowitz SD: Detection of aberrantly methylated hMLH1 promoter DNA in the serum of patients with microsatellite unstable colon cancer. Cancer Res 2001;61:900-902). The analytical sensitivity of the assay is 1%. The tests involves extraction of DNA from FFPE or peripheral blood using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.), PCR amplification and capillary electrophoresis |
Paraffin-Embedded Tissue | Representative formalin-fixed, paraffin-embedded tissue block(s) of tumor tissue is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| MPL Mutation | MPL | Investigation of myeloproliferative neoplasms | University of Alberta Hospital Molecular Pathology | 2 weeks Test performed every second week |
PCR followed by capillary electrophoresis | After DNA extraction, multiplex allele specific PCR amplification for MPL gene exon 10 mutations is performed in two tubes using fluorescently labeled primers to detect the four most common mutations. Amplification products are subjected to fragment analysis by capillary electrophoresis using ABI 3130/3500xL genetic analyzer. Results are analyzed by Genemapper v3.2 software The analytic sensitivity of this assay is 2% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Samples must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| Mucopolysaccharide Screen, urine | MPSCS | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed Monday to Friday Contact for priority turn-around |
Urine | Collect 5 mL (500 uL min) random urine specimen in plain urine container Freeze immediately |
Request using Chemistry Specialty Requisition Very dilute urine samples may give false results and therefore may be cancelled |
||||||||||||||||
| Multiple Endocrine Neoplasia Type 2 | RET | MEN2 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted sequence analysis by NGS | Sequence analysis of exons 10, 11, 13, 14, 15 and 16 Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Multiple Myeloma FISH Panel | CGBM (CO PATH) | University of Alberta Hospital Cytogenetics | 3 weeks from hematopathology report signout Test set-up routinely Monday to Friday |
FISH | Bone Marrow | Collect 2.0 to 4.0 mL (2.0 mL min) of bone marrow into the centrifuge tube with transport media Alternate: Sodium heparin (dark green - NOT PST) |
Specimens should be received in the laboratory within 24 hours of time of collection Specimen MUST remain at room temperature |
Must be accompanied by a completed Hematology Specialty Requisition Ensure all relevant sections of the requisition are complete If multiple myeloma is suspected, this must be indicated on the requistion. List preferred order of FISH probes in the event there are insufficient number of plasma cells for all probes IGH/CCND1 probe testing included Specimens will be rejected if: -Primary tube received refrigerated or frozen -Any aliquot tube is received -Specimen is received in any tube other than sodium heparin -Insufficient specimen volume is received -Specimen received unlabelled -Specimen is received without a complete requisition |
||||||||||||||
| MYC 8q24 by FISH | MYC | Investigation of Burkitt’s lymphoma, Diffuse large B-cell lymphoma | University of Alberta Hospital Molecular Pathology | 1 week for routine or 3 days for RUSH Test set-up on Monday and Wednesday (excluding statutory holidays) |
FISH | Slides are cut from the paraffin block and are deparaffinized in xylene and dehydrated in ethanol. Specimen pretreatment and fluorescence in situ hybridization (FISH) is performed using Vysis® (Abbott Molecular Inc.) LSI MYC Dual Color Break-apart rearrangement probe hybridizes to the band region 8q24. In normal cells, the two probes (green and orange) are fused or close together. In abnormal cells, with a break of 8q24, the two probes separate. This probe detects the translocation: t(8;14)IGH-MYC, which is most characteristic of Burkitt’s lymphoma/leukemia. This probe should also detect two variants t(2;8)IGK-MYC and t(8;22)IGL-MYC. About 5 to 10% of diffuse large B-cell lymphoma (DLBCL) patients also have MYC region rearrangements, and detection of these rearrangements has been associated with a poor prognosis. It has been suggested that FISH analysis for MYC rearrangements should be performed on all DLBCL patients. This lab’s normal cut off is established at <10% of the cells showing an abnormal signal pattern All appropriate negative and positive controls were used. An average of 100 interphase nuclei were examined independently by two observers using an Olympus Provis Fluorescence microscope system on 1000X magnification |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin as well as six 4-micron baked (2 hours at 60°C) unstained charged slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| MYH - Associated Polyposis | SUNQUEST: MYH-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||||
| Myotonic dystrophy type 1 | DMPK | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal, premutation and the majority of expanded pathogenic alleles; very large expanded pathogenic alleles will be detected by repeat-primed PCR but sizing will not be available | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Myotonic dystrophy type 2 | ZNF9 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal and premutation alleles only; detects the presence of expanded pathogenic alleles (no sizing available) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology, Neonatology and Obstetrics Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| N-Acetylaspartic Acid | Investigation of Canavan disease | NAA-U | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with a deuterium-labelled internal standard | N-acetylaspartic acid is significantly elevated in neonatal/infantile (i.e., severe) Canavan disease, but may be only marginally elevated in mild/juvenile Canavan disease | Urine | Collect 5 - 10 mL (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
If specimen is transferred to secondary aliquot tube or container, write "urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided This test should not be ordered in conjunction with urine organic acids, as this metabolite is also reported as part of that test, and this test will be reflexively performed on any urine sample with elevated levels detected Samples will be rejected if: -Aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Nephronophthisis 1 | NPHP1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA | Detects 250kb deletion | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition" |
|||||||||||||
| Neurofibromatosis Type 1 | NF1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Neurofibromatosis Type 2 | NF2 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Newborn Metabolic Screen, blood | Babies born in Alberta are screened for the following disorders: -Amino Acid disorders: PKU, MSUD, CIT, Tyr Ⅰ -Fatty Acid Oxidation disorders: MCAD, LCHAD, VLCAD, CUD, TFP -Organic Acid disorders: MMA, PA, IVA, GA1, HMG, BIOT -Endocrine disorders: CH, CAH -Cystic Fibrosis: CF -Severe Combined Immunodeficiency: SCID -Galactosemia -Sickle Cell Disease: SCD |
NMS | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 4 days Test performed Monday to Saturday |
Blood spot | Newborn Metabolic Screening Blood Spot Requisition (card) | Ensure that expiry date of Newborn Metabolic Screening card (requisition) has not been exceeded before proceeding with collection Collect two saturated blood spots on the Newborn Metabolic Screening card After collection, dry NMS card for a minimum of 3 hours with protective card flap folded back before shipping All information on the Newborn Metabolic Screening card (requisition) must be completed to ensure a valid screen result Ship at ambient temperature Routinely performed on all newborns at 24 hours - 3 days of age. However, if a screen has not been collected in this time frame it is still important that a sample is collected |
For additional information see website www.health.alberta.ca Select tab For Health Professionals, Resources link, Newborn Metabolic Screening link Data Entry: NMS Laboratory enters all requisitions into Specimen Gate as test is not ordered in the LIS Reporting Information: Hard copy reports are distributed to physician offices and the birth facility. Results are available electronically on Alberta Netcare, and most physician office systems |
||||||||||||||
| NPM1 | NPM1 | Investigation of acute myelogenous leukemias (AML) | University of Alberta Hospital Molecular Pathology | 3 business days (5 calendar days) for new AML Test set-up twice a week |
PCR followed by capillary electrophoresis | After DNA extraction, PCR amplification is performed in two separate tubes for FLT3 and NPM1 using fluorescence labeled primers. Amplification products are subjected to fragment analysis by capillary electrophoresis using ABI 3130 genetic analyzer. Results are analyzed by Genemapper v3.2 software The sensitivity of these assays is 5% |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition CEBPA is a reflexed test that is only performed when the assay results for FLT3 / NPM1 are negative and the patient's karyotype is normal |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | Samples must be received in Molecular Pathology within 3 days of collection | ||||||||||||||||||||
| NRAS Mutation Analysis | NRAS | Investigation of colorectal cancers | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Monday and Wednesday (excluding statutory holidays) |
PCR followed by capillary electrophoresis | DNA extraction from FFPE samples is performed using the QIAGEN QIAsymphony DSP DNA mini kit. Analysis for BRAF, KRAS, NRAS mutations is performed after PCR amplification using a single base extension technique (SNaPshot multiplex kit by Life Technologies) and capillary electrophoresis on the 3130 Genetic Analyzer. This multiplexed assay is capable of detecting mutations at BRAF codon 600 and KRAS/NRAS codons 12, 13, 61, 117, and 146. The estimated sensitivity of this assay is 10% |
Paraffin-Embedded Tissue | Representative formalin-fixed, paraffin-embedded tissue block(s) of tumor tissue is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Oculocutaneous Albinism | TYR | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.272G>A (p.C91Y) (NM_000372.4) in the Hutterite population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Ophthalmology, and Dermatology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Oculopharyngeal Muscular Dystrophy | PABPN1 | OPMD | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal, premutation and the majority of expanded pathogenic alleles | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Oligosaccharides Screen, urine | Oligosaccharidoses disorders | OLIGO | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test performed on Tuesday (weekly) | Urine | Collect 5 - 10 mL (1.0 mL min) of random urine sample in plain urine container Freeze immediately |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
|||||||||||||||
| Organic Acids, Semi-Quantitative | Initial investigation of organic acidurias | OA | Alberta Children's Hospital Biochemical Genetics | 14 days Test is performed on a weekly basis |
Derivatization (i.e., silylation) and oximation using gas chromatography-mass spectrometry (GC-MS) | Metabolites reported (truncated list): lactic acid, glycolic acid, glyoxylic acid, oxalic acid, pyruvic acid, 3-hydroxybutyric acid, 2-oxovaleric acid, methylmalonic acid, 4-hydroxybutyric acid, acetoacetic acid, 2-oxo-3-methylvaleric acid, ethylmalonic acid, 2-oxoisocaproic acid, phenylacetic acid, succinic acid, methylsuccinic acid, fumaric acid, glutaric acid, 3-methylglutaric acid, succinylacetone, mandelic acid, malic acid, adipic acid, pyroglutamic acid (a.k.a., 5-oxoproline), 2-hydroxyglutaric acid, 3-hydroxy-3-methylglutaric acid, 2-oxoglutaric acid, suberic acid, 2-oxoadipic acid, orotic acid, homovanillic acid, sebacic acid, vanillylmandellic acid, 3-hydroxypropionic acid, 3-hydroxyisobutyric acid, 3-hydroxy-2-methylbutyric acid, 3-hydroxyisovaleric acid, isovalerylglycine, butyrylglycine, glyceric acid, uracil, 5-hydroxyhexanoic, thymine, propionylglycine, 3-methylglutaconic acid, tiglylglycine, oxaloacetic acid, 3-hydroxyglutaric acid, N-acetylaspartic acid, 2-hydroxyadipic acid, methylcitric acid, homogentisic acid, vanillactic acid, phenylpropionylglycine, indole-3-pyruvic acid, dodecanedoic acid, suberylglycine | Urine | Collect 5 - 10 mL (2.5 mL min) of random urine sample in plain urine container Alternate: 24 hour urine collection container (NO ADDITIVE) Freeze immediately |
Random Urine Collection: 1. If sample is transferred to a secondary aliquot tube or container, write ""random urine"" on label 2. Freeze 24 hour Urine Collection: Urine specimen MUST be kept FROZEN during entire collection and transport process (no preservative) Instruct patient to ensure to place 24 hour urine collection container into freezer immediately after each void throughout the 24 hour collection period After collection: 1. Thaw 24 urine specimen 2. Measure total volume 3. Enter TVOL on requisiton 4. Send 10 mL aliquot of urine; write ""24hr urine, Tvol #L"" on the label 5. Freeze aliquot |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary container or aliquot tube received thawed -Sample type not written on aliquot tube label This test may also be ordered to rule out the possibility of these disorders in a patient being assessed for the ketogenic diet. Abnormal levels found for certain organic acids present in this profile (i.e., 3-methylglutaconic or 3-methylgutaric acids, homovanillic acid, methylmalonic acid, N-acetylaspartic acid, orotic acid, vanillylmandelic acid) will be reflexively quantitated using a separate stable-isotope dilution method on the same sample, volume permitting |
||||||||||||
| Organic Acids, urine | Follow-up test for positive newborn screen (amino acid, organic acid or fatty acid oxidation disorder) Diagnosis of and monitoring for inborn errors of metabolism (patient history/clincal information is essential for adequate evaluation of test results) Elevated anion gap ORGLC |
University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test is performed on a weekly basis |
Urine | Collect 5.0 mL (4.0 mLis acceptable for pediatric samples) of random urine sample in plain urine container For UAH collection only: Put on ice For ALL other sites: Freeze immediately |
If collected at UAH ship on ice. All other sites, ship frozen | Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Urine samples received at room temperature will be rejected |
|||||||||||||||
| Ornithine Transcarbamylase Deficiency | OTC | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Orotic Acid | Investigation of primary and secondary urea cycle disorders, and pyrimidine synthesis disorders resulting in orotic aciduria Orotic acid can be elevated in the following enzyme deficiencies / disorders: -ornithine transcarbamylase deficiency -argininosuccinate synthetase deficiency (i.e., type I citrullinemia) -argininosuccinate lyase deficiency -arginase (ARG1) deficiency -lysinuric protein intolerance -hyperornithinemia, hyperammonemia, and homocitrullinuria (HHH) syndrome -uridine monophosphate synthase (UMPS) deficiency Other causes of elevated orotic acid: -congenital folate deficiency |
OR-U | Alberta Children's Hospital Biochemical Genetics | 21 days Test is performed on a monthly basis |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with a deuterium-labelled internal standard | Urine | Collect 5 - 10 mL (2.5 mL min) of random urine sample in plain urine container Freeze immediately |
If sample is transferred to a secondary aliquot tube or container, write "random urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided. This test should not be ordered in conjunction with urine organic acids, as this metabolite is also reported as part of that test, and this test will be reflexively performed on any urine sample with elevated levels detected Samples will be rejected if: -Aliquot tube received thawed -Sample type not written on aliquot tube label |
|||||||||||||
| Overgrowth Syndrome Panel | CDKN1C, DIS3L2, EZH2, GPC3, NSD1, PTEN | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Pancreatic Cancer Panel | APC, ATM, BRCA1, BRCA2, CDK4, CDKN2A, EPCAM, MLH1, MSH2, MSH6, PALB2, PMS2, STK11, TP53 | Cancer and Endocrine Next Generation Sequencing | SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Pathology | Up to 16 weeks | NGS and Exon Level Array CGH For PMS2: Sanger Sequencing and MLPA |
NGS: Genomic DNA is sequenced on a NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS aCGH: Deletion / duplication testing is performed for those genes (except CDK4, PMS2) at single exon-level resolution MLPA: Detects deletions and duplications Sanger Sequencing: Sequence analysis of PMS2 exons 11-15 only |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
||||||||||||
| Pancreatitis, hereditary | PRSS1 | SUNQUEST: PRSS-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Dectects c.364C>T, c.86A>T and c.86A>C | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Paraganglioma / Pheochromocytoma Predisposition Panel | FH, MAX, RET, SDHAF2, SDHA, SDHB, SDHC, SDHD, TMEM127, VHL, NF1 | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| PDGFRA for GIST | PDGFRA | When ordered in conjunction with KIT, used for the investigation of Gastrointestinal Stromal Tumours (GIST) | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Wednesday |
PCR and sequence analysis | DNA is extracted using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). The exact area of tumorous tissue used in the assay is identified by histological examination and micro dissected from unstained slides PCR amplification is performed using M13 tagged primers. Bidirectional sequencing of the product was performed using the Big dye terminator kit (Life Technologies) and run on the Applied Biosystems 3130 Genetic Analyzer. The obtained DNA sequence was then analyzed using ABI Seqscape® software and compared to the published sequences in the National Center for Biotechnology Information (NCBI) database at www.ncbi.nlm.nih.gov. (GenBank) Analytical sensitivity is 20% |
Paraffin-Embedded Tissue | Representative formalin-fixed, paraffin-embedded tissue block(s) of tumor tissue is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Pediatric Cancer Panel | ALK, APC, BUB1B, CDKN1C, CEP57, DICER1, PHOX2B, PTCH1, SMARCB1, SUFU, TP53, WT1 | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Peutz-Jeghers Syndrome | STK11 | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE CERNER: MDLBL-;APPEND TEST NAME AND SPECIMEN TYPE |
University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception For UAH, Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Phenylalanine & Tyrosine Monitoring | ONLY for monitoring known patients with a diagnosis of phenylketonuria / hyperphenylalaninemia For the investigation of an undiagnosed patient with a suspicion of an amino acid disorder, the Amino Acids - Plasma (ACH) test should be ordered |
PHE, PHE-DBS | Alberta Children's Hospital Biochemical Genetics | 7 days Test is performed daily |
Amino Acid Analyzer with internal standard | Phenylalanine and tyrosine monitoring allows for the assessment of a patient’s dietary phenylalanine tolerance following diagnosis, as well as ongoing adjustments based on their prescribed therapy to ensure the phenylalanine level remains within an acceptable range. It is recommended that all patients on dietary restriction therapy have a full plasma amino acid profile performed at least once a year, in addition to other micronutrient testing, for evaluation of potential nutritional deficiencies | Blood | Collect 1 x 4.0 mL (0.25 mL plasma min) in sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Pre-labelled Blood Spot Collection Card -Random specimen: plain urine container Freeze |
At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze Phenylalanine and Tyrosine Monitoring test may be collected any day, however if results are required by Friday, specimen MUST reach Biochemical Genetics Laboratory (located at ACH) by 1600hr on Wednesday |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provide Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label -Dried blood spots are less than 2/3 full or have not been collected as described above |
||||||||||||
| 7 days Test is performed weekly |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with deuterium-labelled internal standards | Blood Spot | Self-collection of a sample by a patient using the pre-labelled phenylalanine blood spot card requires previous instruction by an ACH metabolic physician or an ACH metabolic dietician If the patient brings a pre-labelled phenylalanine blood spot card to collection lab, warm a finger or heel before collection and fill a minimum of two circles on the card using a single drop of whole blood (blood must soak through to the back of the card) |
Do NOT smear blood onto the card or apply to both sides of card. If a single drop is insufficient, a second drop can be applied. Do NOT use a capillary tube to transfer blood to the card Allow card to dry for a minimum of 4 hours after collection by clipping or lean card against an object on a clean, flat, non-absorbent surface Keep card away from sunlight and heat Place dry collection card inside a biohazard bag Transport at ambient temperature Phenylalanine and Tyrosine Monitoring test may be collected any day, however if results are required by Friday, specimen MUST reach Biochemical Genetics Laboratory (located at ACH) by 1600hr on Wednesday |
||||||||||||||||||
| Phenylalanine / Tyrosine | BPHTY | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 7 days Test is performed Monday to Friday |
Blood spot | Newborn Metabolic Screening Blood Spot Requisition (card) | Ensure that expiry date of Newborn Metabolic Screening card (requisition) has not been exceeded before proceeding with collection Collect two saturated blood spots on the Newborn Metabolic Screening card Use a heel poke if patient is less than one year old or a finger poke if older Could also collect in a Lithium heparin or Sodium heparin tube at any age and apply whole blood to newborn screening card using a capillary tube. Do not allow the capillary tube to contact the card. Each blood spot should require less than 100uL of blood Allow the newborn metabolic screening card to air dry for a minimum of 3 hours. Write ""ACYLCARNITINE, PROFILE"" on the newborn metabolic screening card Ship at ambient temperature in a biohazard bag |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition |
|||||||||||||||
| Pipecolic Acid | Investigation of peroxisomal biogenesis disorders | PIP-P, PIP-S, PIP-U, PIP-C | Alberta Children's Hospital Biochemical Genetics | 30 days Test is performed on a monthly basis /td> |
Derivatization (i.e., silylation) using a gas chromatography-mass spectrometry (GC-MS) stable isotope dilution (SID) method with a deuterium-labeled internal standard | Pipecolic acid is elevated in peroxisomal biogenesis disorders but not in single peroxisomal enzyme deficiencies; therefore, this test in combination with Very Long-Chain & Branched-Chain Fatty Acids testing is useful in differentiating these two classes of peroxisomal diseases Increased pipecolic acid levels may also be seen in alpha-aminoadipic semialdehyde dehydrogenase deficiency (pyridoxine-dependent epilepsy), hyperlysinemia types 1 and 2, and defects of proline metabolism |
Blood | Collect 1 x 4.0 mL (0.25 mL plasma or serum min) in sodium heparin tube (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) -Random specimen: plain urine container -Sterile plastic screw-cap vial for Cerebrospinal Fluid Freeze |
At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided This test should be ordered in conjunction with Very Long Chain & Branched-Chain Fatty Acids (ACH) Samples will be rejected if: -Primary blood tube not refrigerated -Primary CSF tube(s) not kept on ice -Primary urine container not refrigerated or frozen -Aliquot tube received thawed -Tube type not written on aliquot tube label |
||||||||||||
| Urine | Collect 5 - 10 mL (1 mL min) of random urine sample in plain urine container Freeze immediately |
If sample is transferred to a secondary aliquot tube or container, write "random urine" on label | ||||||||||||||||||||
| Cerebrospinal Fluid | Collect 0.25 mL in sterile plastic screw-cap vial for Cerebrospinal Fluid Preference is to use second CSF collection vial with no visible blood contamination Indicate specimen type on label Freeze |
|||||||||||||||||||||
| Porphyria Variegata | PPOX | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect c.175C>T (p.R59W) (NM_001122764.1) in the South African population | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Prader-Willi Syndrome | PWS | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | 15q11 methylation sensitive MLPA and UPD15 Microsatellites | 15q11 methylation sensitive MLPA: Detects copy number changes and determines methylation status. Does not distinguish between imprinting center pathogenic variants and UPD UPD15 microsatellites: Detects UPD15 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| PRF 1-associated Hematological Disorders / Cancers | PRF1 | SUNQUEST: HCA-; APPEND TEST NAME AND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics, Hematology and Oncology |
|||||||||||||
| Prosigna for Breast Cancer | Determine breast cancer recurrence risk in women with hormone receptor positive, HER2 negative, node negative breast cancers with equivocal clinico-pathologic characteristics | University of Alberta Hospital Molecular Pathology | 1 week Test set-up on Monday and Wednesday (excluding statutory holidays) |
Gene expression profile | Prosigna is a Health Canada approved test that uses a 50 gene expression profile, tumour size and nodal status to identify intrinsic subtype, report a Risk of Recurrence Score (ROR) and a probability of Distant Recurrence (DR). RNA is extracted from FFPE tissue, hybridized to the Prosigna CodeSet in the nCounter prep station and scanned by the nCounter Digital Analyzer | Paraffin-Embedded Tissue | Cut the block in the following order: 1. 1 x 4-micron Hematoxylin & Eosin (H&E) stained slide 2. 6 x 10-micron unbaked unstained charged slides 3. 1 x 4-micron Hematoxylin & Eosin (H&E) stained slide |
The breast biomarker pathologist must circle the tumor area in the first H&E slide In addition, the breast biomarker pathologist needs to ensure that the tumor cellularity is >10% and the surface area is > 4 mm2 Please provide a copy of the corresponding pathology report Aptex or Apex slides are not acceptable Electrostatically charged slides must be used |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Test restricted to breast oncologists and pathologists |
|||||||||||||
| Prothrombin G20210A Mutation | Investigation of thrombophilia | PROM | University of Alberta Hospital Molecular Pathology | 2 weeks Test performed on Thursday |
Real-time PCR | After DNA isolation from peripheral blood sample, polymerase chain reaction is performed using the LightCycler 2.0 instrument and the Factor II G20210A kit purchased from F.Hoffman-La Roche Ltd. (Basel, Switzerland) The assay utilizes fluorogenic target-specific hybridization for genotyping of the amplified Factor II DNA and allows the detection of a single point mutation (G to A at position 20210) of the human Factor II gene. Data analysis is performed using the software purchased from the company |
Blood | Collect 1 x 4.0 mL in EDTA (lavender) tube | Keep sample refrigerated until ready for transport Must be received in Molecular Pathology laboratory within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| PTCH 1-related Holoprosencephaly | PTCH1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| PTEN Hamartoma Syndrome and associated developmental disorders | PTEN | "SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE CERNER: MDLBL-;APPEND TEST NAME AND SPECIMEN TYPE" | University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception For UAH, Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology and Pediatrics Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Pyruvic Acid | Used to distinguish between pyruvate dehydrogenase deficiency and other causes of congenital lactic acidosis. This test has no clinical utility outside of the context of elevated lactate. Furthermore, pediatric patients in acute liver failure have also been shown to have elevated lactate and elevated Lactate:Pyruvate (L:P) ratio irrespective of cause (i.e., values did not distinguish between mitochondrial and other causes) | PYR-W, PYR-C | Alberta Children's Hospital Biochemical Genetics | 3 days Prior arrangement is required before ordering |
Detected spectrophotometrically using a commercially-available kit | When blood lactate is > 5 mmol/L, patient classification based on the L:P ratio shows improved diagnostic accuracy: the sensitivity has been reported to be 96% (95% CI, 77%–99%) with a specificity of 100% (95% CI, 59%–100%) | Blood | Collect 1 x 2 mL (0.5 mL whole blood min) in sodium fluoride (grey) tube Alternate: Sterile plastic screw-cap vial for Cerebrospinal Fluid |
For Blood Sample patient must be fasting for 4 hours (including discontinuation of IV dextrose) and must be at rest for 30 minutes prior to collection. Blood must be drawn WITHOUT a tourniquet This test must be booked at the ACH Biochemical Genetics Lab (403-955-7379) at least 24 hours prior to collection, and coordinated with CLS collection staff (403-955-7244). Off-site collections will not be performed A staff member of the Alberta Children’s Hospital Biochemical Genetics Lab must be present at the time of collection to process the specimen |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided This test must be ordered in conjunction with Lactate - Blood or Lactate - CSF. Both tests (Pyruvic Acid and Lactate) will be performed on the same specimen Test will NOT be performed if patient has NOT had previous evidence of elevated blood or CSF lactate level |
||||||||||||
| Cerebrospinal Fluid | Collect 0.5 mL in sterile plastic screw-cap vial for Cerebrospinal Fluid Preference is to use second CSF collection vial with no visible blood contamination Indicate specimen type on label Freeze |
|||||||||||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Rapid Aneuploidy Detection (RAD) | Aneuploidy Turner Syndrome Edward Syndrome Patau Syndrome Down Syndrome Trisomy 13 Trisomy 18 Trisomy 21 |
SUNQUEST: RAD1-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 2-3 days | QF-PCR | Aneuploidy for chromosomes 13, 18, 21, X and Y | Blood (Peripheral or Cord) | For Patients >20 kg - Collect 2 EDTA tubes each containing 3.0 to 5.0 mL of blood For Patients For Neonates - Collect 1 EDTA containing a minimum of 0.5 mL of blood |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Extracted DNA | 10ug @ >100ng/uL | |||||||||||||||||||||
| SUNQUEST: RAD1-; APPEND SPECIMEN TYPE | Chorionic Villi | Minimum of 15 mg | Submit in a sterile 15 mL or 50 mL polypropylene centrifuge tube or in a plain sterile tube with screw cap A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies Chorionic Villi specimens cannot be received on Fridays |
|||||||||||||||||||
| Amniotic Fluid | Minimum of 25 mL | |||||||||||||||||||||
| SUNQUEST: RAD1-; APPEND SPECIMEN TYPE | 6 weeks | Fresh or Frozen Fetal Tissue | > 20mg in a sterile container with NO fixative |
Preferred sample types: Skin, Spleen, Lung, Thymus, Kidney, Muscle or Umbilical Cord Other samples are acceptable but quality of DNA obtained may not be optimal for this assay Products of Conception (POC’s) or placental tissues must be submitted with a maternal blood sample (5ml EDTA) and completed requisition |
||||||||||||||||||
| Cultured Fetal Cells | Confluent (overgrown) T25 flask (or equivalent) placed in transport media | |||||||||||||||||||||
| Renal Cancer Panel | BAP1, EPCAM, FH, FLCN, MET, MLH1, MSH2, MSH6, PMS2, PTEN, SDHA, SDHB, SDHC, SDHD, TP53, VHL, PMS2 | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS For PMS2 - MLPA and Sanger Sequencing |
NGS: Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR MLPA: Detects deletions and duplications Sanger Sequencing: Sequence analysis of PMS2 exons 11-15 only |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Restrictive Dermopathy, lethal | ZMPSTE24 | Fetal hypokinesia sequence | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted Sequence Analysis | Designed to detect the following pathogenic variants: c.54dupT (NM_005857) in the Mennonite population c.1085dupT (NM_005857) in the Hutterite population and Old Colony Mennonite population |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Retinoblastoma | RB1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA (see RAD) | All tumor testing (tissue specimen) to be performed out of province after pre-approval is obtained from the Genetic Resource Center For Tissue Specimens: For Fresh or Frozen Tissue specimens - Submit > 20mg in a sterile container with NO fixative For Cultured Cells - Confluent (overgrown) T25 flask (or equivalent) placed in transport media |
MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Rett Syndrome | MECP2 | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Sequence Analysis and MLPA | Sequence Analysis: Coding region (MECP2a and MECP3b isoforms) MLPA: Detects deletions and duplications |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Rhabdoid Tumor Predisposition Syndrome | SMARCB1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on an NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Rod Cone Dystrophy | RLBP1 | Cone Rod Dystrophy Fundus Albipunctatus |
CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Sanger sequencing, including targeted sequence analysis | Sanger sequencing: Analysis all of coding regions of RLBP1 Targeted sequence analysis: Designed to detect c.141G>A and c.141+2T>C in the Newfoundland population (NM_000326.4) |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Targeted sequence analysis is restricted to those with Newfoundland ancestry |
|||||||||||||
| Russell Silver Syndrome | RSS | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | UPD7 and 11p15 methylation sensitive MLPA | UPD7: Microsatellite markers at 7p14-p11.1 and at 7q32 11p15 methylation sensitive MLPA: Designed to detect abnormal methylation of the two imprinted domains located within the chromosome 11p15 BWS/RSS gene cluster as well as deletions and duplications within the 11p15 region. This assay will detect UPD11 |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| S-Adenosyl-methionine and S-Adenosyl-homocysteine | Further investigation in a patient with hypermethioninemia and/or homocystinuria to distinguish between several disorders affecting methionine metabolism and the methylation cycle S-adenosylmethionine and S-adenosylhomocysteine can be abnormal in the following enzyme deficiencies / disorders: Disorders with elevated S-adenosylmethionine and S-adenosylhomocysteine -cystathionine beta-synthase deficiency (classical homocystinuria) -S-adenosylhomocysteine hydrolase (SAHH) deficiency -adenosine kinase (ADK) deficiency Disorders with elevated S-adenosylmethionine Only -glycine N-methyltransferase (GNMT) deficiency -mtDNA depletion syndrome Disorders with Low-Normal S-adenosylmethionine -methionine adenosyltransferase (MAT) I/III deficiency -methylenetetra-hydrofolate reductase deficiency (MTHFR) -methionine synthase deficiency -Cobalamin (Cbl) C, D, E, F, G, J and X deficiencies -transcobalamin II deficiency -severe folate or B12 deficiencies Other causes of Elevated S-adenosylmethionine and S-adenosylhomocysteine -chronic kidney disease |
SAM-P | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with a deuterium-labelled internal standard | Blood | Collect 1 x 4.0 mL (1.0 mL whole blood min) in K2 EDTA tube (royal blue top) | Collect following a minimum four-hour fast; or just before next feed for newborns Indicate length of fast, or time of last feeding on the requisition (with collection time) Place sample immediately on ice Transport immediately to ACH Biochemical Genetics Laboratory or immediately call ACH Biochemical Genetics Laboratory (403-955-7379) for sample pick-up Should collection be required at a site other than ACH, contact the ACH Biochemical Genetics Lab PRIOR TO sample collection for instructions on how to properly handle the sample |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided |
|||||||||||||
| Schwannomatosis Panel | NF2, SMARCB1 | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Shwachman-Diamond Syndrome | SBDS | SUNQUEST: MDG-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | Up to 16 weeks | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on an NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS aCGH: Deletion/duplication testing is performed at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics, Hematology, and Oncology |
|||||||||||||
| Sialic Acid - Total & Free | Investigation of neuraminidase deficiency (including galactosialidosis), lysosomal free sialic acid storage disorders and sialuria. This test may also be used in the investigation of I-cell disease (resulting from a functional neuraminidase deficiency) Disorders detectable by elevated Total and Free Sialic Acid: -lysosomal free sialic acid disorders (i.e., Salla disease, intermediate severe Salla disease, and infantile free sialic acid storage disease) -sialuria Disorders detectable by elevated Total and Conjugated Sialic Acid: -sialidosis (a.k.a., neuraminidase deficiency) -galactosialidosis (a.k.a., cathepsin A deficiency) -I-cell disease (severe phenotype) Other causes of elevated Total, Free and/or Conjugated Sialic Acid: -hemolytic uremic syndrome -pregnancy -type 2 diabetes mellitus -renal failure -bladder tumors |
SIALIC-U | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis |
Liquid chromatography, electrospray-ionization tandem mass spectrometry (LS-MS/MS) with a deuterium-labelled internal standard | Urine | Collect 5 - 10 mL (0.5 mL min) of random urine sample in plain urine container Freeze immediately |
If sample is transferred to a secondary aliquot tube or container, write "random urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Aliquot tube received thawed -Sample type not written on aliquot tube label |
|||||||||||||
| Skeletal Dysplasias Panel - hypochondroplasia, achondroplasia, thanatophoric | FGFR3 | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | Targeted sequence analysis | Sanger sequencing of exons 3, 5-9, 11-12, 14-15 and 18 in FGFR3. Designed to detect the following common pathogenic variants (NM_000142.4): Achondroplasia c.1138G>A (p.Gly380Arg) c.1138G>C (p.Gly380Arg) Hypochondroplasia c.1620C>A (p.Asn540Lys) c.1620C>G (p.Asn540Lys) Thanatophoric dysplasia, type 1 c.742C>T (p.Arg248Cys) c.746C>G (p.Ser249Cys) c.1108G>T (p.Gly370Cys) c.1111A>T (p.Ser371Cys) c.1118A>G (p.Tyr373Cys) c.1949A>T (p.Lys650Met) c.2420G>T (p.Ter807LeuextTer101) c.2419T>G (p.Ter807GlyextTer101) c.2419T>C (p.Ter807ArgextTer101) c.2419T>A (p.Ter807ArgextTer101) c.2421A>T (p.Ter807CysextTer101) c.2421A>C (p.Ter807CysextTer101) c.2421A>G (p.Ter807TrpextTer101) Thanatophoric dysplasia, type 2 c. 1948A>G (p.Lys650Glu) |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics, Perinatology and Pediatrics |
|||||||||||||
| Skin Cancer Panel | BAP1, CDK4, CDKN2A, CYLD, PTCH1, SUFU | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Sphingomyelinase, T-lymphocyte | Investigation of Niemann-Pick A & B Disease (a.k.a., sphingomyelinase deficiency) - a lysosomal storage disorder | SPHINGO | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis | Sphingomyelinase activity is measured spectrophotometrically using the 2-N-Hexadecanoylamino-4-nitrophenyl-phosphoroylcholine-linked substrate | This procedure involves the establishment of an IL-2 stimulated T-lymphocyte culture and storage of cell pellets for the purposes of lysosomal enzyme testing | Blood | Collect 1 x 4.0 mL (3.0 mL whole blood min) in sodium heparin (dark green - NOT PST) Alternate: Lithium heparin (dark green - NOT PST) |
Primary tube must remain at room temperature at all times It is recommended that patients go to ACH OP Lab for specimen collection The 3.0 mL of whole blood will be sufficient to test for multiple lysosomal enzymes in T-lymphocytes, if requested If not collected at ACH, all samples SHOULD arrive before 14:00h and have been collected NO LONGER than 48 hours prior to arrival |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Requests for testing should be supported by a high index of clinical suspicion specific to the disorder in question Samples will be rejected if: -Primary tube not kept at room temperature -Any aliquot tube is received. -Any sample other than whole blood is submitted |
||||||||||||
| Spinal and Bulbar Muscular Atrophy | AR | SBMA Kennedy Disease |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal and premutation alleles only; detects the presence of expanded pathogenic alleles (no sizing available) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Spinal Muscular Atrophy | SMN1, SMN2 | Arthrogryposis Multiplex Congentiaspinal Muscular Atrophy Kugelberg Welander Syndrome Werndig Hoffman Syndrome SMA |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA | Detects dosage (including deletions and duplications) in exons 7 & 8 of the SMN1 and SMN2 genes | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Hemoglobinopathy screen results are required |
|||||||||||||
| Spinocerebellar Ataxia General Screen | ATXN1 (SCA1), ATXN2 (SCA2), ATXN3 (SCA3), CACNA1A (SCA6), ATXN7 (SCA7), ATXN8 (SCA8) | Machado-Joseph Disease Spinocerebellar Ataxia Type 1 Spinocerebellar Ataxia Type 2 Spinocerebellar Ataxia Type 3 Spinocerebellar Ataxia Type 6 Spinocerebellar Ataxia Type 7 Spinocerebellar Ataxia Type 8 SCA SCA screen |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal and premutation alleles only; detects the presence of expanded pathogenic alleles (no sizing available) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Each SCA type can also be ordered individually based on clinical suspicion or family history. On the requistion, specify if request is for general screen or specific SCA type Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Spinocerebellar Ataxia Type 17 | TBP | SCA SCA17 |
CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR sizing and repeat-primed PCR | Detects and sizes normal and premutation alleles only; detects the presence of expanded pathogenic alleles (no sizing available) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
||||||||||||
| Steroid Sulfatase Deficiency | NLGN4X , FAM16AX, STS, KAL1, OA1 | X-linked Ichthyosis STS Deficiency |
SUNQUEST: STS-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA + Real-Time PCR | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Sugar (reducing substances) Screen | Initial investigation for disorders of galactose and fructose metabolism | REDSUB-U | Alberta Children's Hospital Biochemical Genetics | 8 days Test is performed on a weekly basis |
Benedict’s test | Positive and negative controls are run with patient samples | Urine | Collect 5 - 10 mL (0.5 mL min) of random urine sample in plain urine container Freeze immediately |
If sample is transferred to a secondary aliquot tube or container, write "random urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test should not be used to monitor known patients on treatment as this is a qualitative test A positive Benedict’s test indicating the presence of increased urine reducing substances will result in reflexive sugar chromatography testing on the same sample, volume permitting Samples will be rejected if: -Primary container or aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Sugar Chromatography | Used to evaluate issues of carbohydrate absorption, when there is a strong clinical suspicion of a disorder of monosaccharide transport or carbohydrate metabolism (e.g., galactosemia, fructosemia), or when the sugar (i.e., reducing substances) screen has been found to be abnormal and there was insufficient sample for reflexive testing | SUGAR | Alberta Children's Hospital Biochemical Genetics | 21 days Test is performed on a biweekly basis |
Thin layer chromatography with known sugar standards | Metabolite reported: rhaminose, xylose, fucose, glucose, fructose, galactose, sucrose, lactose, raffinose | Urine | Collect 5 - 10 mL (1 mL min) of random urine sample in plain urine container Freeze immediately |
If sample is transferred to a secondary aliquot tube or container, write "random urine" on label If sample is transferred to a secondary aliquot tube or container, write "random urine" on label | Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided Samples will be rejected if: -Primary container or aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Sulfite Screen | Investigation of patients suspected of having isolated sulfite oxidase or molybdenum co-factor deficiencies based on clinical presentation | SULFITE | Alberta Children's Hospital Biochemical Genetics | 1 day Prior arrangement is required before ordering |
Colourometric test strip | Although a clearly positive test is highly suggestive of either disorder, it is dependent on testing fresh urine A negative test cannot be used to rule out these disorders, and further testing is recommended if warranted by specific clinical suspicion |
Urine | Collect 5 - 10 mL (2.0 mL min) of random urine sample in plain urine container Place sample on ice |
This test must be booked with the ACH Biochemical Genetics Lab (403-955-7379) perferably 24 hours in advance Sample can only be collected at ACH Immediately transport to ACH Biochemical Genetics Lab or call ACH Biochemical Genetics Lab (403-955-7379) for sample pick-up It is recommended that testing be performed on a freshly voided sample, preferrably within 20 minutes of collection. As sulfite rapidly oxidizes at room temperature as well as durring refrigeration, testing will be cancelled if the sample has not been received within 1 hour of collection |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
| Synovial Sarcoma by FISH | SS18 | Investigation of Synovial Sarcoma | University of Alberta Hospital Molecular Pathology | 2 weeks Test set-up on Monday and Wednesday (excluding statutory holidays) |
FISH | Slides are cut from paraffin blocks and are deparaffinized in xylene and dehydrated in ethanol. Specimen pretreatment and fluorescence in situ hybridization (FISH) are performed using Vysis® (Abbott Molecular Inc.) pretreatment kit The locus specific identifier (LSI) SS18 (SYT) dual colour break apart gene rearrangement probe is a mixture of two FISH DNA probes that hybridize to opposite sides of the SS18 gene. Synovial sarcoma is characterized by a t(X;18)(p11.2;q11.2) which is present in virtually all examples. This translocation fuses SYT on chromosome 18 with either SSX1 (66%) or SSX2 (33%), both located on chromosome Xp11. More rarely, SYT is adjoined to SSX4 also located on chromosome Xp11. The majority of breakpoints for t(X;18)(p11.2;q11.2) is not found at random and arises exclusively in Synovial sarcoma. In normal cells, the two probes (green and orange) are fused or close together. Normal nuclei contain two red/green (yellow) fusion signals (2F). In abnormal cells, with t(X;18)(p11.2;q11.2), having a breakpoint within the hybridization targets of the probes exhibits one yellow, one red and one green signal (FRG) This lab’s normal cut off is established at less than or equal to 10% of the cells showing an abnormal signal pattern. All appropriate negative and positive controls are used An average of 100 interphase nuclei are examined independently by two observers using an Olympus Provis Fluorescence microscope system on 1000X magnification |
Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin as well as six 4-micron baked (2 hours at 60°C) unstained charged slides can be submitted |
Please provide a copy of the corresponding pathology report | MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| t(15;17) PML/RARA | PML, RARA | Investigation of myeloproliferative neoplasms and/or acute leukemias | University of Alberta Hospital Molecular Pathology | Detection Assay - RUSH 1 day, routine 3 days Quantitative Assay - 1 week Test performed Monday to Friday |
Real-time PCR | PML/RARA Detection Assay: RNA extraction is performed on the QIAsymphony. cDNA synthesis and amplification of t(15;17) gene transcripts is performed in two steps by RT qPCR with ABL1 as the reference gene. Fluorescence detection is by the Quant Studio real time PCR platform (Life Technologies). Appropriate positive and negative controls are used The limit of detection (LOD) of this assay is a 5.0 log reduction from the mean of pooled positive diagnostic samples PML/RARA Quantitation Assay: RNA extraction is performed on the QIAsymphony. cDNA synthesis and quantitation of t(15:17) gene transcripts is performed in two steps by RT-qPCR with ABL1 as the reference gene. Fluorescence detection is by the Quant Studio real time PCR platform (Life Technologies). Appropriate positive and negative controls are used The results are expressed as percentage residual disease and a calculated log reduction (LR). The residual disease is the percentage of NCN of the monitor sample to the NCN of the initial diagnostic sample. The LR is the log reduction in the normalized copy number (NCN) of PML-RARA transcript from the initial diagnostic sample. The NCN is the ratio of PML-RARA transcripts to ABL1 transcripts The limit of detection (LOD) of this assay is a 5.0-log reduction from the mean of pooled positive diagnostic samples, which corresponds to 0.001% residual disease |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology Collect 1 x 2.5 mL PAXgene Blood RNA for Molecular Pathology |
Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Using a butterfly needle and holding the tube below the arm to prevent backflow of contents into the patient, first collect 2 EDTA tubes, then the PAXgene Blood RNA tube Collection must be in the correct order of draw to ensure accurate white count determination For bone marrow collection, order of draw is not important Immediately following collection, place all tubes ON ICE Transport samples to Molecular Pathology as soon as possible, on ice or cold pack Molecular Pathology must receive the sample within 24 hours of collection If transport to Molecular Pathology is anticipated to be greater than 24 hours, contact Molecular Pathology at 780-407-6648 |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition PAXgene collection tubes are available upon request. Submit an order form using one of the links provided: Client Supply Order Edmonton Zone OR Client Supply Order Provincial Form |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube Collect 1 x 2 mL (max) PAXgene Blood RNA tube |
|||||||||||||||||||||
| t(8;21) RUNX1/RUNX1T1 | RUNX1, RUNX1T1 | Investigation of myeloproliferative neoplasms and/or acute leukemias | University of Alberta Hospital Molecular Pathology | 1 week Test performed Monday to Friday |
Real-time PCR | RNA extraction is performed on the QIAsymphony. cDNA synthesis and quantitation of t(8:21) gene transcripts is performed in a single well using the Express One-Step Superscript qRT PCR mastermix (Life Technologies) with ABL1 as the reference gene. Fluorescence detection is performed on the Quant Studio real time PCR platform (Life Technologies). Appropriate positive and negative controls are used The results are expressed as percentage residual disease and a calculated log reduction (LR). The residual disease is the percentage of NCN of the monitor sample to the NCN of the initial diagnostic sample. The LR is the log reduction in the normalized copy number (NCN) of RUNX1/RUNX1T1 transcript from the initial diagnostic sample. The NCN is the ratio of RUNX1/RUNX1T1 transcripts to ABL1 transcripts The limit of detection (LOD) of this assay is a 4.0-log reduction from the mean of pooled positive diagnostic samples, which corresponds to 0.01% residual disease |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology Collect 1 x 2.5 mL PAXgene Blood RNA for Molecular Pathology |
Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Using a butterfly needle and holding the tube below the arm to prevent backflow of contents into the patient, first collect 2 EDTA tubes, then the PAXgene Blood RNA tube Collection must be in the correct order of draw to ensure accurate white count determination For bone marrow collection, order of draw is not important Immediately following collection, place all tubes ON ICE Transport samples to Molecular Pathology as soon as possible, on ice or cold pack. Molecular Pathology must receive the sample within 24 hours of collection If transport to Molecular Pathology is anticipated to be greater than 24 hours, contact Molecular Pathology at 780-407-6648 |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition PAXgene collection tubes are available upon request. Submit an order form using one of the links provided: Client Supply Order Edmonton Zone OR Client Supply Order Provincial Form |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube Collect 1 x 2 mL (max) PAXgene Blood RNA tube |
|||||||||||||||||||||
| t(9;22) BCR/ABL1 | BCR, ABL1 | Investigation of myeloproliferative neoplasms and/or acute leukemias | University of Alberta Hospital Molecular Pathology | Qualitative assay: 1) AML and ALL: 3 business days (5 calendar days) 2) Other samples: 8 business days (10 calendar days) Quantitative assay: 1 month for monitor cases Initial cases are set-up every other day (not including weekends or statutory holidays) Monitor cases are set-up biweekly, on Wednesday |
Detection assay - PCR followed by capillary electrophoresis. Quantitation assay - Real-time PCR |
BCR/ABL1 Detection Assay: RNA extracted from the sample is reversed transcribed and the resultant cDNA is subjected to PCR. This qualitative, lab-developed assay is designed as a multiplex PCR. The amplified DNA is then run on the 3500xL genetic analyzer for detection and analysis The assay is capable of detecting nine different BCR/ABL1 fusion transcripts: b2a2, b3a2, e1a2 (most commonly found in CML and ALL) as well as b2a3, b3a3, e6a2, e8a2, e1a3 and e19a2 BCR amplification is also measured in every sample to test whether a sample has sufficient quantity and quality of RNA The limit of detection (analytical sensitivity) for b2a2, b3a2 and e1a2 fusions was determined to be 3.5 log reduction p210 BCR/ABL1 Quantitation Assay: After RNA isolation and cDNA synthesis, real-time quantitation of BCR-ABL1 gene transcripts was performed using the BCR-ABL Mbcr IS-MMR Kit purchased from QIAGEN (Hilden, Germany). Appropriate positive and negative controls were used Results are reported on the International Scale (IS) which is calibrated against the first NIBSC WHO IS standard (ref. 09/138) The analytical sensitivity of the UAH Molecular Pathology assay is 0.0032% (4.5 Log Reduction) |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology Collect 1 x 2.5 mL PAXgene Blood RNA for Molecular Pathology |
Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Using a butterfly needle and holding the tube below the arm to prevent backflow of contents into the patient, first collect 2 EDTA tubes, then the PAXgene Blood RNA tube Collection must be in the correct order of draw to ensure accurate white count determination For bone marrow collection, order of draw is not important Immediately following collection, place all tubes ON ICE Transport samples to Molecular Pathology as soon as possible, on ice or cold pack. Molecular Pathology must receive the sample within 24 hours of collection. If transport to Molecular Pathology is anticipated to be greater than 24 hours, contact Molecular Pathology at 780-407-6648 |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition PAXgene collection tubes are available upon request. Submit an order form using one of the links provided: Client Supply Order Edmonton Zone OR Client Supply Order Provincial Form |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube Collect 1 x 2 mL (max) PAXgene Blood RNA tube |
|||||||||||||||||||||
| Tay-Sachs CARRIER Testing | NOT appropriate for patients with a suspected diagnosis of Tay-Sachs or Sandhoff disease. For diagnostic testing for these disorders, please see Hexosaminidase (ACH) | TAYSACHS | Alberta Children's Hospital Biochemical Genetics | 30 days Monthly, with priority testing being performed on samples from pregnant females |
Measured fluorometrically using 4-methylumbelliferyl substrates | Total hexosaminidase, as well as fractional and specific hexosaminidase A are assessed using activities in both serum and lymphocytes, and these activities are compared to established reference ranges for noncarriers, Tay Sachs carriers, Sandhoff carriers and pregnant females. Either serum or lymphocyte total hexosaminidase activities that fall outside of the established ranges cannot be used to determine carrier status using that specimen type. Activities that fall within a known overlap between Tay-Sachs carrier and noncarrier (i.e., the indeterminate) reference ranges also will not allow determination of carrier status | Blood | Collect 2 x 4 mL (3 mL serum min) Non-additive (red top - NOT SST) tubes Collect 3 x 4 mL (7 mL whole blood min) Sodium heparin (dark green - NOT PST) tubes |
This test must be booked with the ACH Biochemical Genetics Lab (403-955-7379) at least 24 hours in advance Collections are recommended to be done at the ACH Outpatient Lab between 8:00 and 14:00 weekdays, as a member of the ACH Biochemical Genetics Lab will be required to retrieve and prepare the sample Serum Sample: 1. Centrifuge 2. Transfer serum to an aliquot tube 3. Write ""Red Top"" on the aliquot tube label 4. Freeze Serum sample can be refrigerated in primary tube (following clotting) prior to freezing in an aliquot tube Whole Blood Sample: Primary tube must remain at room temperature at all times If samples are collected at ACH: Immediately transport to ACH Biochemical Genetics Lab or call ACH Biochemical Genetics Lab (403-955-7379) for sample pick-up If samples are not collected at ACH: Whole blood sample must be transported at room temperature to arrive in ACH by 1400hr and within 24 hours of time of collection, as lymphocyte quality (isolated from the green top tubes) drops after 24 hours and/or if not maintained at room temperature |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Physicians only: Please indicate on requisition if patient is Jewish, Ashkenazi Jewish, pregnant, diabetic, on birth control or on estrogen therapy Order Restrictions: This test will not be performed on minors Testing requires serum AND whole blood sample collections from BOTH partners (e.g. husband AND wife) and will ONLY be conducted through a physician referral with appropriate genetic counseling services |
||||||||||||
| T Cell Receptor Gene Rearrangement | TCR | Investigation of lymphoproliferative disorders TCR Gene Rearrangement | University of Alberta Hospital Molecular Pathology | TCR gamma 7 days, TCR beta 14 days Test set-up on Monday and Thursday (excluding statutory holidays) |
PCR followed by capillary electrophoresis | DNA is extracted using the QIAsymphony DSP DNA Mini Kit from Qiagen (Valencia, CA.). The integrity of the DNA is assessed using a Specimen Control Size Ladder detecting four different fragment sizes. Appropriate polyclonal and monoclonal (T-cell lymphoma cell line) controls are used. A no template control is also included. These assays have been validated by a parallel study with another institution T-Cell Receptor Gamma Assay: A four-color, multiplex, PCR assay is performed using Invitrogen Platinum High Fidelity Taq with appropriate fluorescence-labeled primers detecting four different fragment sizes. The PCR products are subjected to capillary electrophoresis on an Applied Biosystems 3130/3500XL Genetic Analyzer using fragment analysis GeneMapper software. Previously published primers for the four-color TCR gamma PCR Assay (Vega et al, Am J Clin Pathol 2001;116:17-24) are utilized in this method. Due to the sensitivity of this assay, pseudoclonality can be seen occasionally in some inflammatory conditions. These clones are usually not reproducible when a repeat sample is obtained. This assay cannot detect less than 5 positive cells/95 normal cells The clinical sensitivity of this assay is determined to be 87.5% and clinical specificity is 95% in our laboratory T-Cell Receptor Beta Assay: A commercially available kit from InVivoscribe Technologies (San Diego, California) is used. PCR is performed using Invitrogen Platinum High Fidelity Taq with appropriate fluorescence-labeled primers in three different mixes. The PCR products are subjected to capillary electrophoresis on an Applied Biosystems 3130/3500XL Genetic Analyzer using fragment analysis GeneMapper software. This assay does not detect 100% clonal cell populations and cannot detect less than 5 positive cells/95 normal cells The clinical sensitivity of this assay is determined to be 52% and clinical specificity is 84% in our laboratory |
Blood | Collect 2 x 4 mL EDTA (lavender) tubes. One tube for CBC/differential and one tube for Molecular Pathology | Any peripheral blood requests will require a CBC and differential to be ordered and processed as per local handling practices Whole blood samples must be received in Molecular Pathology within 3 days of collection |
MUST be submitted on a Molecular Pathology Requisition Please provide any pertinent clinical history on the requisition Test requests are restricted to hematopathologists and lymphoma pathologists |
||||||||||||
| Bone Marrow | Collect 1 x 4 mL (2 mL min) EDTA (lavender) tube | |||||||||||||||||||||
| Paraffin-Embedded Tissue | A formalin-fixed, paraffin-embedded tissue block is preferred Alternatively, one slide stained with Hematoxylin & Eosin and five 10-micron non-baked, unstained slides can be submitted |
Please provide a copy of the corresponding pathology report | ||||||||||||||||||||
| Telomere MLPA | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Torsion Dystonia, Autosomal Dominant | TOR1A | DYT1 | CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR and Targeted Sequence Analysis | Designed to detect c.907_909delGAG (NM_000113) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Neurology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Tuberous Sclerosis Panel | TSC1, TSC2 | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Urine Sugar Identification | Pentosuria and Fructosuria | SUGID | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 10 days Test is performed Monday to Friday | Urine | Collect 1 mL (100 uL min) of random urine sample in plain urine container Freeze immediately |
First morning void urine sample is preferred | Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Does not detect sucrose If sucrase intestinal deficiency is suspected, order disaccharidase panel (requires small bowel biopsy sample) If diabetes is suspected, order random or fasting glucose If galactosemia is suspected, order galactosemia screen. If pediatric intestinal lactase deficiency is suspected, order fecal sugar identification |
||||||||||||||
| Urine Sulfite | Sulfite oxidase deficiency and molybdenum cofactor defieciency | MISC-; URINE SULFITE | University of Alberta Hospital Newborn Metabolic Screen & Biochemical Genetics | 4 days Test is performed Monday to Friday |
Urine | Collect 1 mL of random urine sample in plain urine container Freeze immediately |
Collect Monday to Friday before 1400 hours (excluding holidays) only Notify Newborn Metabolic Screening / Biochemical Genetics Laboratory (780) 407-8820 when patient presents at collection site, and before collection Keep sample on ice Samples must be received in the Newborn Metabolic Screening / Biochemical Genetics Lab on ice within 2 hours of collection Test ideally performed within 1 hour of collection |
Request using Chemistry Specialty Requisition Please ensure all required information is provided on the requisition Rejection Criteria: Samples not received in lab, on ice within 2 hours of collection If baby is on TPN, this will cause a false positive result |
||||||||||||||
| Uronic Acids, Quantitative | Strong clinical suspicion of a mucopolysaccharidosis in a patient, or when the initial glycosaminoglycan (i.e., mucopolysaccharide) screen was found to be abnormal and there was insufficient sample for reflexive testing | UA-U | Alberta Children's Hospital Biochemical Genetics | 21 days Test performed on a biweekly basis |
Determined spectrophotometrically following precipitation with a cationic detergent, washing and hydrolysis with carbazole reagent | Elevated uronic acids indicate the presence of abnormal levels of the glycosaminoglycans dermatan and/or heparan sufate in addition to the normal urine constituents chondroitin 4 and 6 sulfates, and is highly suggestive of a biochemical diagnosis of a mucopolysaccharidosis. Such a finding requires further identification of which glycosaminoglycan(s) has contributed to this finding. Further enzyme or molecular-based testing is required to confirm the specific enzyme deficiency in each case NOTE: this method does not detect keratan sulfate, as neither of its constituent subunits (i.e., galactose and N-acetylglucosamine) is a hexuronic acid. Therefore, in patients with a strong clinical suspicion of Morquio syndrome, both quantitative uronic acids AND GAG electrophoresis should be ordered |
Urine | Collect 5 - 10 mL (4 mL min) of random urine sample in plain urine container Alternate: 24 hour urine collection (10.0 mL min) container (NO ADDITIVE) Freeze immediately |
Random Urine Collection: 1. Prefer first morning void whenever possible 2. If sample is transferred to a secondary aliquot tube or container, write ""random urine"" on label 3. Freeze 24 hour Urine Collection: Urine specimen MUST be kept FROZEN during entire collection and transport process (no preservative) Instruct patient to ensure to place 24 hour urine collection container into freezer immediately after each void throughout the 24 hour collection period After collection: 1. Thaw 24 urine specimen 2. Measure total volume 3. Enter TVOL on requisiton 4. Send 10 mL aliquot of urine; write ""24hr urine, Tvol #L"" on the label 5. Freeze aliquot |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition This test is considered specialty testing; this test will be cancelled if sufficient clinical justification is not provided This test can either be ordered in conjunction with Glycosaminoglycan Electrophoresis, Qualitative (ACH) , or, if the level of uronic acids is found to be elevated in a urine sample, Glycosaminoglycan Electrophoresis, Qualitative will be reflexively tested and reported separately. Minimum volume is for both tests Samples will be rejected if: -Aliquot tube received thawed -Sample type not written on aliquot tube label |
||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Very Long-Chain & Branched-Chain Fatty Acids | Investigation of peroxisomal biogenesis disorders and specific peroxisomal fatty-acid oxidation disorders Disorders Detectable by very long-chain and branched-chain fatty acid analysis*: Peroxisomal Biogenesis Disorders -Zellweger spectrum disorder (encompassing Zellweger syndrome, NALD and infantile Refsum disease) Specific Peroxisomal Enzyme Deficiencies: -X-linked adrenoleukodystrophy -acyl-CoA oxidase deficiency (a.k.a., pseudo-NALD) -D-bifunctional protein deficiency -2-methylacyl-CoA racemase deficiency -Adult Refsum disease *Further confirmatory testing is required for some of these conditions, as nonspecific/artifactual findings as well as overlapping patterns may prevent definitive diagnosis in specific cases |
VLCFA-P, VLCFA-S | Alberta Children's Hospital Biochemical Genetics | 45 days Test is performed on a monthly basis | Derivatization (i.e., silylation) using a gas chromatography-mass spectrometry (GC-MS) stable isotope dilution (SID) method with deuterium-labeled internal standards | Metabolite reported: behenic (docosanoic) acid (C22:0), lignoceric (tetracosanoic) acid (C24:0), ceratinic (hexacosanoic) acid (C26:0), pristanic (2,6,10,14-tetramethylpentadecanoic) acid, phytanic (3,7,11,15-tetramethylhexadecanoic) acid | Blood | Collect 1 x 4.0 mL (0.25 mL plasma or serum min) in sodium heparin (dark green - NOT PST) Alternate: -Lithium heparin (dark green - NOT PST) -Non-additive (red top - NOT SST) |
Collect following a minimum four-hour fast; or just before next feed for newborns Patients should avoid eating peanut-rich goods (e.g., peanut butter) 24 hours prior to collection Indicate length of fast, or time of last feeding on the requisition (with collection time) At ACH only: The primary heparinized tube can be put on ice (ensure not to freeze sample) or refrigerated for routine pick-ups between 8:30 am – 2:30 pm, Monday through Friday For all other collections: Centrifuge, transfer plasma or serum to aliquot tube, write ""NaHep"", ""LiHep"" or ""Red Top"" on the aliquot tube label and freeze |
Request using Biochemical Genetics Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Samples will be rejected if: -Primary tube not refrigerated -Aliquot tube received thawed -Tube type not written on aliquot tube label This test should be ordered in combination with plasma/serum or urine pipecolic acid for the investigation of a peroxisomal biogenesis disorders as an additional, independent measure of peroxisomal function This test SHOULD NOT be ordered for the investigation of very long chain acyl-CoA dehydrogenase deficiency, as the enzyme deficiency in that disorder is involved in mitochondrial (not peroxisomal) fatty acid beta-oxidation For the investigation of very long chain acyl-CoA dehydrogenase deficiency, order Acylcarnitine Profile (ACH) |
||||||||||||
| von Hippel-Lindau Syndrome | VHL | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | MLPA and Sanger sequencing | Sanger Sequencing: Coding region of VHL gene MLPA: Detects partial or whole gene deletions and duplications |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Confirmation of Diagnosis (patient has signs or symptoms of the disease/disorder) - Genetics, Endocrinology Presymptomatic Testing (patient does not presently have symptoms, positive family history of the condition) - Genetics |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Werner Syndrome | WRN | SUNQUEST: MDG-; APPEND TEST NAME AND SPECIMEN TYPE /td> | University of Alberta Hospital Molecular Diagnostics | NGS and Exon Level Array CGH | NGS: Genomic DNA is sequenced on an NGS instrument. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS aCGH: Deletion/duplication testing is performed at single exon-level resolution |
Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Williams-Beuren (7q11.2) Locus (deletion and duplication syndromes) | Infantile Hypercalcemia Elfin Faces with Hypercalcemia Supravalver Aortic Stenosis William Syndrome |
SUNQUEST: WMS-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Real-Time PCR | Detects deletions and duplications | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
|||||||||||||
| WT1-Related Disorders | WT1 | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
|||||||||||||
| Test Name | Gene(s) | Alternate Test Name/ Indication | Test Code | Performing Site | TAT | Method | Detection details | Specimen Requirements | Specimen Handling | Requisition Form/ Restrictions | ||||||||||||
| Sample | Process | |||||||||||||||||||||
| Xeroderma Pigmentosum Panel | DDB2, ERCC2, ERCC3, ERCC4, ERCC5, XPA, XPC | Cancer and Endocrine Next Generation Sequencing | CERNER: MDLBL-; APPEND TEST NAME AND SAMPLE TYPE | Alberta Children's Hospital Molecular Diagnostics | Up to 16 weeks | NGS | Genomic DNA is sequenced on a NGS instrument. NGS detects nucleotide substitutions, small insertions and deletions, and copy number variants. Sanger sequencing is used to confirm all variants with clinical or uncertain significance and to analyze regions with <90% technical sensitivity by NGS. Additional deletion/duplication testing may be performed by a variety of methods, including, but not limited to: multiplex ligation-dependent probe amplification and quantitative PCR | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception | MUST be submitted on a Molecular Diagnostic Laboratory Cancer and Endocrine Next Generation Sequencing Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition Order Restrictions: Genetics |
||||||||||||
| X-Inactivation studies | SUNQUEST: MDL-; APPEND TEST NAME AND SPECIMEN TYPE CERNER: MDLBL-; APPEND TEST NAME AND SPECIMEN TYPE |
University of Alberta Hospital or Alberta Children's Hospital Molecular Diagnostics | 6 - 8 weeks | PCR | Androgen receptor CAG analysis (requires heterozygosity at this locus) | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception For UAH, Chorionic Villi specimens cannot be received on Fridays |
For MDL North (UAH), MUST be submitted on a Molecular Diagnostic Laboratory Requisition For MDL South (ACH), MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||||
| Y Chromosome Microdeletion | AZF | Chromosome Y Deletion AZF Azoospermia Male Infertility Y Deletion |
SUNQUEST: DELY-; APPEND SPECIMEN TYPE | University of Alberta Hospital Molecular Diagnostics | 6 - 8 weeks | Microsatellites | 7 Y chromosome loci | Blood, DNA, Prenatal and Fetal Tissue (see RAD) | A maternal blood sample (5ml EDTA) must be collected for maternal cell contamination studies for products of conception Chorionic Villi specimens cannot be received on Fridays |
MUST be submitted on a Molecular Diagnostic Laboratory Requisition Please ensure all required information, including patient's clinical history/indication, is provided on the requisition |
||||||||||||
This article has been produced by Merogenomics Inc. and edited by Jason Chouinard, B.Sc. Reproduction and reuse of any portion of this content requires Merogenomics Inc. permission and source acknowledgment. It is your responsibility to obtain additional permissions from the third party owners that might be cited by Merogenomics Inc. Merogenomics Inc. disclaims any responsibility for any use you make of content owned by third parties without their permission.
Products and Services Promoted by Merogenomics Inc.
Select target group for DNA testing
Healthy screening |
Undiagnosed diseases |
Cancer |
Prenatal |
Or select popular DNA test
 |
 |
 |
 |
Pharmaco-genetic gene panel |
Non-invasive prenatal screening |
Cancer predisposition gene panel |
Full genome |