COVID-19 and genetic predisposition
Dr.M.Raszek
To have symptoms or no symptoms? That is the question!
Everyone in the world probably knows by now that if someone gets infected by SARS-CoV-2 virus, the resulting impact can range from a complete lack of symptoms to severe respiratory failure leading to death.
The accumulating data so far show that the highest risk factor for succumbing to a severe case of COVID-19 is first and foremost old age, with other notable risk factors being overweight or obese (and other associated comorbidities that are also frequently reported in general), as well as being male. But these factors do not help at all to explain the broad spectrum of clinical symptoms that people of nearly all age groups can experience once badly infected with the coronavirus.
Obviously, one possible contributing factor that would be worth investigating is the role of our personal genetics. We previously touched lightly on this topic when looking at potential COVID-19 susceptibility by ethnicity due to differences in the expression of the ACE2 receptor that SARS-CoV-2 virus uses to enter human cells. That was at the start of pandemic and since then, genetic susceptibility to COVID-19 has harnessed lots of attention as so many have perished due to this SARS-CoV-2 pandemic.
So, what have we learned so far?
Related to Neanderthals? Not just embarrassing, but even deadly
One very interesting report that came out late last year is based on the work of the COVID-19 Host Genetics Initiative which compared the genetics of those who were hospitalized due to the SARS-CoV-2 infection versus an unaffected control population. This particular study employed the genome-wide association study (GWAS) technique, which is a method frequently discussed in the Merogenomics blog - it is where you compare two different population groups and attempt to identify potential genetic differences between them. In just this way one variant (a mutation) on chromosome 3 was identified that stood out between COVID-19 affected patients and controls. The ID of the variant is rs35044562 for those of you who have had your genome sequenced and want to find out. Although be careful what you seek! Or are you brave enough to find out what lurks beneath your hood?
What is interesting about the location of this variant is that it happens to reside inside a haplotype (a genomic segment inherited from one generation to another in one chunk) that is inherited from Neanderthals! That’s right, COVID-19 predisposition inherited from Neanderthals!
We explained the concept of haplotypes when we wrote about a unique client’s case of discovering hearing loss predisposition due to the inheritance of a specific group of variants that were always found grouped together in affected individuals. In essence, inheriting DNA from your parents, you get it through a random shuffle of large genomic chunks called haplotypes. When you pass your genetics to your offspring, you will also pass it on through this random assortment of the genetics you inherited from your parents. Thus, as you can imagine, the closer the ancestor, the larger the size of haplotypes are inherited. The older the ancestor, the smaller the inherited haplotype size because there are more generations between you and the ancestor for this random process of shuffling genetics to take place as haplotypes are being passed from parents to offspring. In the case of Neanderthals, we are talking about ancestors that were 40 to 60 thousand years ago!
Of the 13 variants found to be present in that haplotype, 11 of them are also found in the genome of the approximately 50,000-year-old Vindija 33.19 Neanderthal from modern day Croatia. But only three of these variants are observed in the genomes of older Neanderthals from Siberia.
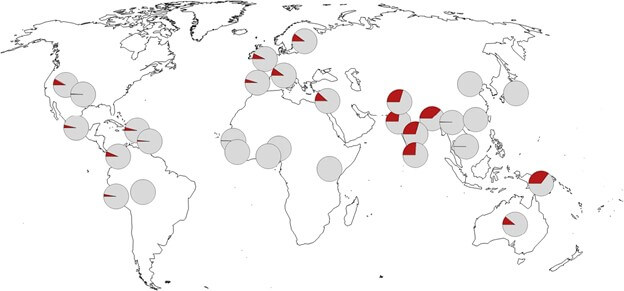
In terms of the distribution of this Neanderthal-inherited haplotype among the global population, around 50% of people in South Asia carry at least one copy of the risk haplotype, 16% of people in Europe, 9% of the people in the Americas but it is almost completely absent in populations from Africa.
What is interesting is that people in east Asia are also almost completely devoid of having this haplotype. The authors speculated that either the inheritance of haplotype was selected for elsewhere in the world for some protective impact, or perhaps there was negative selection pressure against it in east Asia due to other past coronavirus-like outbreaks. Which means right now the rest of the world is experiencing a similar negative selection against this haplotype due to the COVID-19 pandemic by increasing the risk of dying and not passing on the haplotype to next generation.
That is basically how evolution works, folks. A random game of genetic cards where the same genetic material can confer advantages in one millennium, and destroy the population in another.
How your immune system should work!
Another more recent finding for genetic predisposition to COVID-19 was related to the quality of personal immune systems. That probably will not sound too surprising but we especially wanted to dive into this topic just to review for our readers a particular and important component of immune function.
When a virus like SARS-CoV-2 infects a cell, it hijacks the cell’s mechanisms to direct the cell to make many copies of the virus. Since pictures are worth a thousand words, check out this short Merogenomics video on the topic below.
But some of our cells have the ability to break down viruses, and present their fragments on the cell surface. It is these fragmented pieces of virus that are then recognized by a specific type of T-cells, called cytotoxic T-cells or killer T-cells, and once they identify a virus fragment, these T-cells make copies of themselves to produce an army that will attack any similar invading virus and kill them. In a matter of few days, a killer T-cell cavalry is mounted against the invaders.
But the focus of this tale is actually not the killer T-cells, as cool as they are. It is the molecules directly involved in the process by which those cleaved virus components get to the surface of the cells. For example, viral proteins like the now famous, or infamous, spike protein that crowns the virus and is used to bind to ACE2 receptors and then enter human cells. It is this very spike protein, by the way, that is heavily mutating in those new emerging variants (in a bid to escape our immune attack against it.) These cut-up protein fragments in general are called peptides, and we covered this concept in one of our past posts dedicated to… the genetics of love. Trust us, check it out, as it will blow your mind.
But back to our story!
The biological system of molecules that is involved in presenting virus protein peptide fragments for recognition by T-cells is the human leukocyte antigen (HLA) class I molecules. You have likely heard about this before, but who can remember every little detail in this complex biological myriad, right? Well these molecules are important to remember as they are the internal receptors that bind virus fragments to present them on the surface of the cell.
There are three main types of HLA class I receptors: HLA-A, HLA-B, and HLA-C. Everybody inherits one different copy from each of our parents. Overall, there are many variations of HLA-A, HLA-B, and HLA-C in the population (these are also referred to as variants), and each one of them has its own distinct ability to recognize various foreign virus proteins.
Genetic lottery for your viral luck
And now is where that genetic variation’s predisposition comes in. Because these HLA class I receptors are so variable, the distribution of specific types is driven by evolution, and so what types of pathogen a given population has to combat with, and hence each population, or each country if you prefer, will have its own specific variants present. It is already well known that what type of HLA class I receptors a person has will predispose them to how severe a viral infection will be, and thus it should not be surprising that the same effect would be observed with the SARS-CoV-2 viral infection. Basically, it is the chance that whether you have the distribution of highly protective HLA class I receptors that can effortlessly recognize cleaved SARS-CoV-2 viral proteins and present them to mount a killer T-cells response, or not.
This is exactly what some researchers have demonstrated when they analyzed HLA class I receptors variants in relation to the severity of COVID-19. However, this was only computationally demonstrated, with different SARS-CoV-2 virus peptides modelled against different HLA class I receptors found in the studied population. But in this way, they collected affinity values between 6,548 SARS-CoV-2 peptides and 107 HLA class I variants. In this way, the best and worst HLA class I receptors were identified in terms their ability to present SARS-CoV-2 peptides. Then the distribution of these receptors was looked at in the COVID-19 deceased patients, and mapped against the age of death. In this way, it was found that certain HLA class I variants were associated with an increased risk of death in people under the age of 60 as compared to controls or others of low risk.
Further, it is interesting that regarding the HLA class I variants linked to high-risk, people who had two such copies (one inherited from each of their parents – referred to as being homozygous for such variant), were at an even greater risk of dying. Conversely, those who were homozygous for low-risk HLA class I variants were even less likely to die when infected. So, the authors called it a double-edge sword. Being homozygous for a specific HLA class I variant could massively protect you against one type of virus, while it could kill you when infected with another type.
Again, for those who have means of investigating this (at your own risk), HLA-A*02:01 and HLA-A*03:01 were associated with low risk while HLA-A*01:01 was associated with the high-risk group. This, in fact, has also already been observed and published before with regards to Italian patients’ susceptibility to COVID-19, further giving weight to this finding.
Genetic COVID-19 predisposition, digging deep
In this last section we will focus on a publication presented to Merogenomics by one of our past clients! The study was performed by Genetics Of Mortality In Critical Care out of UK, once again using a GWAS approach to compare COVID-19 critically ill patients with controls from population genetic studies (the UK is one of the world leaders in linking the use of genomics to personal health, and they have developed some of the most comprehensive, mouth-watering databases in the world). The authors found a number of variants which they then analyzed across other available genetic data with information related to COVID-19. By the way, that analysis included 23andMe data from a respiratory study, documenting which clients were placed on a ventilator, had oxygen administered or had pneumonia.
In this way the authors identified a number of variants associated with COVID-19 severity: rs10735079 on chromosome 12 in a gene cluster (OAS1, OAS2 and OAS3) that is responsible for development of activators of antiviral molecules that cut virus genomes; rs74956615 on chromosome 19 near the TYK2; rs2109069 also on chromosome 19 within the DPP9 gene; and rs2236757 on chromosome 21 in the IFNAR2 gene.
These authors studied the expression of these genes, meaning the use of the gene as a blueprint to produce RNA (which itself is a blueprint) to produce a tiny molecular protein robot that performs some function within the cellular universe. Yes, the same category or type of RNA that is delivered in the new mRNA vaccines, which explains how they work, by providing a tiny blueprint on how to build some viral spike protein inside our own cells, which are then presented on the surface of those cells to activate the immune system. If COVID-19 vaccines are a topic of interest to you, check out Merogenomics YouTube videos dedicated to the topic.
When authors studied the expression of these genes, they found that low expression of IFNAR2 (meaning the gene use for production of RNA and subsequent protein is supressed) was associated with life-threatening COVID-19. This gene encodes a receptor for interferon, which is a molecule that informs production of many components of the early immune response to viral infection. Such molecules involved in stimulating immune responses and inflammation are referred to as cytokines. It would make sense you would want to see more of these receptors available.
On the other hand, high expression of TYK2 and CCR2 was associated with severe COVID-19. TYK2 encodes a protein involved in stimulation of numerous cytokines. CCR2 protein is also a receptor that once activated promotes migration of certain white blood cells towards a site of inflammation. This can be good but then also bad if overdone which can be seen in patients in the acute phase of COVID-19 with inflammatory lung injury. Thus, these two genes could be involved in promoting such injury if overused.
Finally, DPP9 gene encodes a protease, which is a protein that cleaves something else, usually to promote a specific biological reaction. In this case, this protease can also trigger a certain immune response, inflammation, possibly acting as a certain checkpoint in activating these events.
Oh yeah, they also identified a variant on chromosome 3 that has previously been reported in other studies as linked to severe COVID-19: rs73064425. Well, remember that Neanderthal haplotype with a number of variants travelling together in time through inheritance? This appears to be one of those variants, found inside that Neanderthal COVID-19 susceptibility haplotype genetic region! Poor Neanderthals just cannot catch a break! Kidding! Deeper analysis of that same region by another group also found that within that haplotype there also might be some protective variants, and so more careful analysis is needed when determining which ethnicities might be more or less in danger due to this genetic region.
What to do with all that genetics
There is constantly new information coming out related to genetic susceptibility to COVID-19 and here is our selection of some that we thought would be the most interesting. If you had your genome decoded, you could look for specific information but be wary that just because associations have been found and published, it is still a far cry from being clinically valid. It would require careful observation of many more DNA sequences of those affected and unaffected by COVID-19 to be able to confirm such findings. If you elect to investigate these, keep that in mind that such results might not be entirely accurate in predicting disease severity.
In addition, even if the presenting genetic information is contributing to disease development, like any personal risk to disease development based on genetics, there can always be other mitigating factors - both within your genetics as well as outside your body - that could be mitigating your individual ability to produce diseases and their clinical symptoms. This holds true for all genetic investigations. Just because pathogenic mutations are uncovered it never fully guarantees that a condition will materialize, and therefore that should always be kept in mind: be prudent and take smart mitigating actions, but do not be consumed by the fear of something that might never come to pass.
This is a reason why DNA sequencing is not for everyone. But, if it is for you, educate yourself and select your DNA tests wisely (the Merogenomics site is a great resource!) There is lots of garbage out there, and if you come across a DNA test claiming they can inform you of your personal risk of COVID-19, well, take it as educational, but with a grain of salt.
The same caution goes if you wish to investigate your already decoded genome.
This article has been produced by Merogenomics Inc. and edited by Jason Chouinard, B.Sc. Reproduction and reuse of any portion of this content requires Merogenomics Inc. permission and source acknowledgment. It is your responsibility to obtain additional permissions from the third party owners that might be cited by Merogenomics Inc. Merogenomics Inc. disclaims any responsibility for any use you make of content owned by third parties without their permission.
Products and Services Promoted by Merogenomics Inc.
Select target group for DNA testing
Healthy screening |
Undiagnosed diseases |
Cancer |
Prenatal |
Or select popular DNA test
 |
 |
 |
 |
Pharmaco-genetic gene panel |
Non-invasive prenatal screening |
Cancer predisposition gene panel |
Full genome |